The common symbiosis signaling pathway (CSSP) is a signaling cascade in plants that allows them to interact with symbiotic microbes. It corresponds to an ancestral pathway that plants use to interact with arbuscular mycorrhizal fungi (AMF). It is known as "common" because different evolutionary younger symbioses also use this pathway, notably the root nodule symbiosis with nitrogen-fixing rhizobia bacteria. The pathway is activated by both Nod-factor perception (for nodule forming rhizobia), as well as by Myc-factor perception that are released from AMF. The pathway is distinguished from the pathogen recognition pathways, but may have some common receptors involved in both pathogen recognition as well as CSSP. A recent work [1] by Kevin Cope and colleagues showed that ectomycorrhizae (a different type of mycorrhizae) also uses CSSP components such as Myc-factor recognition.
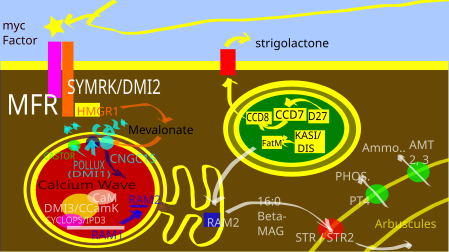

The AMF colonization requires the following chain[2] of events that can be roughly divided into the following steps:
1: Pre-Contact Signaling
2: The CSSP
2: A: Perception
2: B: Transmission
2: C: Transcription
3: The Accommodation program
Outline
editTo accurately recognize the infection thread of a different species of organism, and to establish a mutually beneficial association requires robust signaling.[3] AM fungi are also fatty acid auxotrophs;[4][2] therefore they depend on a plant for supply of fatty acids.[5]
At the pre-symbiotic signaling, plants and AMF release chemical factors in their surroundings so that the partners can recognise and find each other.[6]' Plant root exudates play roles in complex microbial interaction,[7] by releasing a variety of compounds,[7][8][9] among which strigolactone has been identified to facilitate both AMF colonisation and pathogen infection.[8]
Phosphate starvation in plant induces strigolactone production as well as AMF colonisation.[8] Plants release strigolactone, a class of caroteinoid-based plant hormone, which also attracts the fungal symbionts, and stimulate the fungal oxidative metabolism along with growth and branching of the fungal partner. [2] Strigolactone promotes hyphal branching in germinating AMF spores and facilitates colonisation. [9] [10]
The common symbiosis signalling pathway is called so because it has common components for fungal symbiosis as well as rhizobial symbiosis. The common signalling pathway probably evolved when the existing pathway for arbuscular mycorrhizae was exploited by rhizobia.[2][11]
The perception happens when fungal Myc factor is detected by the plant. Myc factors are comparable to rhizobial nod factors. The chemical nature of some Myc-factors has recently been revealed as lipo-chito-oligosaccharide (Myc-LCOs) and chito-oligosaccharides (Myc-COs) that work as symbiotic signal.[10][12][13]
The presence of Strigolactone enhances the production of Myc-CO production by AMF.[10]
Myc-factor receptor (MFR) is still putative. However, a protein called DMI2 (or SYMRK) has a prominent role in perception process and it is thought to be a co-receptor of MFR. Other plants such as rice may employ different mechanisms using OsCERK1 and OsCEBiP to detect chitin oligomers.[2][14][15] However, recent work has demonstrated that rice SYMRK is essential for AM symbiosis.[16]
The transmission happens when the signal is transmitted after detection to the plant nucleus. This process is mediated by two nucleoporins NUP85 and NUP133,[11] Alternatively, another hypothesis says HMG-CoA reductase is activated on perception, which then converts HMG-CoA into mevalonate. This mevalonate acts as a second messenger and activates a nuclear K+ cation channel (DMI-1 or Pollux).[2][17] The transmission stage ends by creating a ‘calcium spike’ in the nucleus. [18]
The transcription stage starts when a Calcium and Calmodulin dependent kinase (CCaMK) is activated.[2] This kinase stimulates a target protein CYCLOPS.[2] CCaMK and CYCLOPS probably forms a complex that along with DELLA protein, regulates the gene expression of RAM1 (Reduced Arbuscular Mycorrhyza1).[2]
The accommodation process involves the extensive remodelling of host cortical cells. This includes invagination of host plasmalemma, proliferation of endoplasmic reticulum, golgi apparatus, trans-golgi network and secretory vesicles. Plastids multiply and form “stromules”. Vacuoles also undergoe extensive reorganization.[11]
The Pre-Contact Signaling
editChemical signalling starts prior to the two symbionts coming into contact. From the host plant's side, it synthesizes and releases a range of caroteinoid based phytohormone, called strigolactones.[2] They have a conserved tricyclic lactone structure also known as ABC rings.[19] Strigolactone biosynthesis occurs mainly in plastid,[20] where D27 (Rice DWARF 27; Arabidopsis ortholog ATD27), an Iron binding beta-carotene isomerase works at upstream of strigolactone biosynthesis. [20] Then carotenoid cleavage dioxygenase enzyme CCD7 and CCD8 modifies the structure, which has following orthologs:
| Gene name | Localization | function | Rice ortholog | Pea ortholog | Petunia ortholog | Arabidopsis ortholog |
|---|---|---|---|---|---|---|
| CCD7 | Plastid proteins | involved in strigolactone biosynthesis | D17/ HTD1 | RMS5 | DAD3 | MAX3 |
| CCD8 | Plastid proteins | involved in strigolactone biosynthesis | D10 | RMS1 | DAD4 | MAX4 |
| Alpha/Beta fold hydrolase | Nuclear proteins | involved in strigolactone perception | D14 | RMS3 | DAD2 | ? |
The alpha/beta fold hydrolase D3 and also D14L (D14-Like) (the later one has an Arabidopsis ortholog, KAI2, or KARRIKIN INSENSITIVE-2) is reported to have important roles in mycorrhizal symbiosis,[21] notably, D3, D14 and D14L are localised in the nucleus.[2]
NOPE1 or 'NO PERCEPTION 1', is a transporter protein in Rice (Oryza sativa) and Maize (Zea mays), also required for the priming stage for colonisation by the fungus. NOPE1 is a member of Major Facilitator Super family of transport proteins, capable of N-acetylglucosamine transport. Since nope1 mutants root exudates fail to elicited transcriptional responses in fungi, it strongly seems that NOPE1 secretes something (not yet characterised) that promotes fungal response.[22]
Perception
editThere are two main type of root symbiosis; one is root nodule symbiosis by Rhizobia (RN-type) and another is Arbuscular Mycorrhiza (AM-type). There are common genes involved in between these two pathways.[23] these key common components, form the Common Symbiosis pathway (CSP or CSSP).[23] It has been proposed that, RN symbiosis has originated from AM symbiosis.[11] The perception of the presence of the fungal symbiont takes place mainly through fungal chemical secretions generally termed as Myc-factors. Receptors for Myc-factors are yet to be identified. However, DMI2/SYMRK probably acts as a co-receptor of Myc factor receptor (MFR). The AM fungal secreted materials relevant to symbiosis are Myc-LCOs, Myc-COs, N-Acetylglucosamine [2][24]
| Myc factor | Plant protein it mainly act on |
|---|---|
| Myc-LCOs | LYS11 in Lotus japonicus |
| Short chain chitin oligomers (COs) | OsCERK1 and OsCEBiP in rice |
| N-acetylglucosamine | NOPE-1 in maize |
Fungal Molecules that triggers CSSP
editMyc-LCOs (lipochitooligosaccharides)
editLike Rhizobial LCOs (Nod factors); Myc-LCOs play important role in perception stage. They are a kind of secreted compounds from AM fungi, mainly mixtures of lipo-chito-oligosaccharides (Myc-LCOs). In Lotus japonicus, LYS11, a receptor for LCOs, was expressed in root cortex cells associated with intra-radical colonizing arbuscular mycorrhizal fungi [24]
Short chain chitin oligomers (Myc-COs)
editAM host plants show symbiotic-activated calcium waves upon exposure to short chain chitin oligomers. It has been reported that production of these molecules by the AM fungus Rhizophagus irregularis, is strongly stimulated upon exposure to strigolactones. [2] This suggests that plants secrete strigolactones and in response, the fungus increases short chain chitin oligomers, which in turns elicits the plant response to accommodate the fungus. The lysine motif domain of OsCERK1 and OsCEBiP is thought to be involved in the perception of short chain chitin oligomers.[2]
NOPE-1 is transporter (described above). NOPE-1 also shows a strong N-acetylglucosamine uptake activity, and is thought to be associated with recognition of presence of fungal symbiont.[2]
Some plant proteins are suspected to recognise Myc-factors, and the rice OsCERK1 Lysin motif (LysM) receptor-like kinase, is one of them.[15]
Cell Surface Receptors
editThere are multiple families of pattern recognition receptors and co-receptors involved in recognition of microbial pathogens and symbionts. Some of the relevant families involved in CSSP, are Membrane bound LysMs (LYM), Soluble LysM Receptor like Protein, LYK (LysM receptors with active Kinase domain), LYR (LysM proteins with inactive kinase domain), etc.
Seemingly, different combinations of a LYK and LYR receptors perceive and generate differential signals, such as some combinations generate a pathogen recognition signal whereas some combinations generate symbiotic signals. [25][26][27][28]
Receptor-like Kinases (RLKs)
editDMI2/ SYMRK is a receptor-like kinase, an important protein in endosymbiosis signal perception, reported in several plants (Mt-DMI2 or Mt-NORK in Medicago trancatula; Lj-SYMRK in Lotus japonicas; Ps-SYM19 in Pisum sativum; OsSYMRK in Rice). OsSYMRK lacks an N-terminal domain and exclusively regulate AM symbiosis (is not involved in the RN symbiosis).[29] Notably, it has been found that a Nod-factor inducible gene, MtENOD11, is activated in the presence of AMF exudates; little is known about this phenomenon.[30][31]
LysM receptor-like kinase
editLysin Motif (LysM) receptor-like kinase are a subfamily related to membrane bound Receptor-like kinase (RLKs) with an extracellular region consisting of 3 Lysine motifs. They have some important orthologs in different plants, that vary in their function. In some plant species they are involved in AM symbiosis, in others they are not. Tomato (Solanum lycopersicum), a non-legume eudicot, also have a similar LysM receptor, SlLYK10 that Promotes AM symbiosis. There are some co-receptors of Myc-factor receptor viz., OsCEBiP in Rice, a LysM membrane protein can function as a co-receptor of OsCERK1 but it participates in a different pathway.[32][33][34]
Most of these kinases are serine/threonine kinases, some are tyrosine kinases.[27] Also, they are type-1 transmembrane proteins, that indicates their N-terminal domain towards the outside of the cell, and the C-terminal domain is towards inside of the cell.[27]
| Medicago truncatula | Lotus japonicus | Pisum sativum
(pea) |
Prunus persica | Arabidopsis thalliana | Brassica rapa | Solanum lycopersicum
(Tomato) |
Brachypodium distachyon | Oryza sativa
(Rice) | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lysine Motif
Receptor-Like Kinase and Lysine Motif Receptor like Protein |
LYM | LYMI | LYM1 | PpLYM1 | AtLYM1
AtLYM3 |
SlLYM1 | BdLYM1
BdLYM3 |
OsLYP6
OsLYP5, OsLYP4 | |||||
| LYMII | LYM2 | PpLYM3
PpLYM2 |
AtLYM2 | SlLYM3
SlLYM2 |
BdLYM2
BdLYM4 |
OsCEBiP
OsLYP3 | |||||||
| LYR | LYR 1 | LYRIA | MtNFP
MtLYR1 |
LjNFR5
LjLYS11 |
PpLYR1 | SlLyk10 | Bd LYR1 | OsNFR5 | |||||
| LYRIB | MtLYR8 | PpLYR2 | SlLYK9 | Bd LYR2 | |||||||||
| LYR 2 | LIRIIA | MtLYR10 | LjLYS16 | PpLYR6 | AtLYK2 | SlLYK2 | |||||||
| LYRIIB | MtLYR9 | LjLYS15 | PpLYR7 | SlLYK15 | |||||||||
| LYR 3 | LYRIIIA | MtLYR3 | LjLYS12 | PpLYR3 | AtLYK4 | SlLYK4 | Bd LYR4 | OsLYK6 | |||||
| LYRIIIB | MtLYR2 | PpLYR4 | SlLYK7
SlLYK6 |
||||||||||
| LYRIIIC | MtLYR4
MtLYR7 |
LjLYS13
LjLYS14 |
AtLYK5 | Bd LYR3 | OsLYK3
OsLYK2, OsLYK4 | ||||||||
| LYR 4 | LYRIV | MtLYR5
MtLYR6 |
LjLYS20 | PpLYR5 | |||||||||
| LYK | LYKI | LYK1, LYK4, LYK5, LYK6, LYK7, LYK2, LYK3, LYK9, LYK8 | LjLYS2
LjLYS1, LjNFR1, LjLYS6, LjLYS7 |
PpLYK2
PpLyk1 |
AtLYK1/
AtCERK1 |
SlLYK13
SlLYK1/ SlBti9, SlLYK12, SlLYK11 |
BdLYK1 | OsCERK1 | |||||
| LYKII | LYK10 | LjLYS3/
EPR3 |
PpLYK3
PpLYK4 |
||||||||||
| LYKII | PpLYK5 | AtLYK3 | SlLYK3 | BdLYK3 | |||||||||
| Receptor like Kinase | RLK | Mt-DMI2/
Mt-NORK |
Lj-SYMRK | Ps-SYM19 | OsSYMRK | ||||||||
Transmission
editThe transmission of signal cascades into the nucleus is not well understood. However, this transmission includes carrying the message up to the nuclear membrane and generation of a calcium wave.[35] Some elements involved in this process are:
Nucleoporins
editLotus japonicus Nucleoporins LjNUP85 and LjNUP133 has potential role in transmission of the signal.[36] Lj-NENA is another important nucleoporin that plays role in AM symbiosis.[29]
HMGR and Mevalonate.
editIt has been proposed that the enzyme 3-hydroxy-3-methylglutaryl-CoA reductase (HMG CoA reductase or HMGR) has potential role in the transmission stage. The enzyme is activated by SYMRK/DMI2, and forms mevalonate.[37][38] This mevalonate acts as a second messenger, and activates a nuclear potassium channel, DMI1 or POLLUX.[38]
| Nuclear envelope Protein | Function | Rice | Lotus japonicus | Medicago truncatula | Pisum |
| CNGC15 | Cyclic-nucleotide gated Calcium-channel | Mt-CNGC15 | |||
| Castor | Potassium cation channel | Os-Castor | Lj-Castor | ||
| POLLUX or DMI1 | Potassium cation channel | OsPOLLUX | LjPOLLUX | Mt-DMI1 | Ps-SYM8 |
Nuclear membrane cation channels.
editThe nuclear calcium channel CNGC15, which is cyclic nucleotide gated ion channel; mediates the symbiotic nuclear Ca2+ influx, and it is countered by K+ efflux by DMI1.[37]
Transcription
edit| Protein | Function | Name of the Plant | |||
| Rice | Lotus japonicus | Medicago Truncatula | Pisum sativum | ||
| CCamK | Calcium calmodulin-dependent kinase with role in AMF symbiosis | Os-DMI3 or
Os-CCaMK |
Lj-CCaMK | Mt-DMI3 | Ps-SYM9 |
| CYCLOPS | Coiled coil domain containing proteins that respond to CCamK and promote AMF symbiosis | Os-CYCLOPS | Lj-CYCLOPS | Mt-IPD3 | Ps-SYM33 |
| DELLA | Promote AMF symbiosis | Os-SLR1 | Mt-DELLA1
Mt-DELLA2 |
Ps-LA
Ps-CRY | |
Calmodulin is a widespread regulatory protein that functions along with Ca2+ in various biological processes. In AM symbiosis signalling, it modulates CCaMK.[37] CCaMK or DMI3 is a calcium-and-calmodulin-dependent kinase (CCaMK) thought to be a key decoder of Ca2+ oscillations and an important regulatory kinase protein. Nuclear Ca2+ spiking promotes binding of Ca2+ calmodulin with CCaMK.[37] Binding of Ca2+ calmodulin with CCaMK causes conformational change of CCaMK that stimulates a target protein, CYCLOPS, which has different orthologs.[37] CYCLOPS is a coiled coil domain containing protein [37] possibly form a complex with CCaMK[37] that works along with DELLA proteins. DELLA proteins are a kind of GRAS-domain protein originally identified as repressors of the Gibberellin signalling pathway, however now it is seen that DELLA participates in many signalling pathways.[40] There are two DELLA proteins in Medicago trancatula and Pisum sativum that play a role in symbiosis whereas in rice only one DELLA protein fulfils this task.[37] Reduced Arbuscular Mycorrhiza or RAM1[37] is a GRAS[41] protein whose gene is directly regulated by DELLA and CCaMK/ CYCLOPS.[37] By using chromatin immunoprecipitation assays, it has been shown that RAM1 binds to RAM2 gene promoter.[37] RAM1 also regulates many of the plant genes that participate in AMF accommodation.
Some GRAS proteins play a role in AM symbiosis but these roles are not yet fully understood. These include RAM1, RAD1 (REQUIRED FOR ARBUSCLE DEVELOPMENT 1), MIG1 (MYCORRHIZA INDUCED GRAS1), NSP1 and NSP2.[42] WRKY transcription factor genes are thought to play very important roles in establishment of mycorrhizal symbiosis and they perhaps work through regulating plant defense genes.[43]
The Accommodation program
editRoot cortex cells experience important changes in order to accommodate for the fungal endosymbiont. The pre-penetration apparatus (PPA) in outer cell layers and the peri-arbuscular membrane that surrounds arbuscules in inner cell layers need to be formed and the plant cell cytoplasm needs to rearrange,[36] the vacuole retracts in size, the nucleus and nucleolus enlarge in size and chromatin decondense indicating heightened transcriptional activity.[36] Plastids multiply and stay connected with “stromulus”.[36] Furthermore, it was suggested that the apoplastic longitudinal hyphal growth is probably regulated by plant genes such as taci1 and CDPK1.[44]
Genes and proteins playing a role in the accommodation programme
editAlthough various proteins have been identified which may play role on how this accommodation process occurs, the detailed signalling cascade is not fully understood. Some of the proteins and mechanisms involved in the deposition on peri-arbuscular membrane are EXOCYST complex, EXO70 subunit, a symbiosis-specific splice variant of SYP132, VAPYRIN, and two variants of VAMP721.[37] Plant enzymes FatM and RAM2[45] and ABC transporter STR/STR2 are putatively involved in the synthesis and supplying of a lipid 16:0 β-monoacylglycerol to the AM fungi.[45][46] Recently discovered kinases that regulate the AMF accommodation programm include ADK1,[47] AMK8, AMK24,[48] ARK1[49] and ARK2.[50]
The protein composition of the peri-arbuscular membrane is very different from that of the plasma membrane. It includes some special transporters such as phosphate transporters (Mt-PT4, Os-PT11, Os-PT13) and ammonium transporters (Mt-AMT2 and 3). It also includes ABC transporters such as STR/STR2 putatively involved in lipid transport.[37][51]
Evolutionary significance
editAM fungi and plants co-evolved and developed a very complex interaction that allow the plant accommodate the AM-fungal host.[52][53][54] It has been proposed that the RN symbiosis has originated from the AM symbiosis.[36][29]
See also
editReferences
edit- ^ Cope, Kevin R.; Bascaules, Adeline; Irving, Thomas B.; Venkateshwaran, Muthusubramanian; Maeda, Junko; Garcia, Kevin; Rush, Tomás A.; Ma, Cathleen; Labbé, Jessy; Jawdy, Sara; Steigerwald, Edward; Setzke, Jonathan; Fung, Emmeline; Schnell, Kimberly G.; Wang, Yunqian; Schleif, Nathaniel; Bücking, Heike; Strauss, Steven H.; Maillet, Fabienne; Jargeat, Patricia; Bécard, Guillaume; Puech-Pagès, Virginie; Ané, Jean-Michel (October 2019). "The Ectomycorrhizal Fungus Laccaria bicolor Produces Lipochitooligosaccharides and Uses the Common Symbiosis Pathway to Colonize Populus Roots". The Plant Cell. 31 (10): 2386–2410. doi:10.1105/tpc.18.00676. PMC 6790088. PMID 31416823.
- ^ a b c d e f g h i j k l m n o MacLean, Allyson M.; Bravo, Armando; Harrison, Maria J. (October 2017). "Plant Signaling and Metabolic Pathways Enabling Arbuscular Mycorrhizal Symbiosis". The Plant Cell. 29 (10): 2319–2335. doi:10.1105/tpc.17.00555. PMC 5940448. PMID 28855333.
- ^ Barman, Jagnaseni; Samanta, Aveek; Saha, Babita; Datta, Siraj (December 2016). "Mycorrhiza: The oldest association between plant and fungi". Resonance. 21 (12): 1093–1104. doi:10.1007/s12045-016-0421-6. ISSN 0971-8044. S2CID 109932925.
- ^ Luginbuehl, Leonie H.; Menard, Guillaume N.; Kurup, Smita; Van Erp, Harrie; Radhakrishnan, Guru V.; Breakspear, Andrew; Oldroyd, Giles E. D.; Eastmond, Peter J. (16 June 2017). "Fatty acids in arbuscular mycorrhizal fungi are synthesized by the host plant". Science. 356 (6343): 1175–1178. doi:10.1126/science.aan0081. PMID 28596311. S2CID 206658467.
- ^ Bravo, Armando; Brands, Mathias; Wewer, Vera; Dörmann, Peter; Harrison, Maria J. (June 2017). "Arbuscular mycorrhiza-specific enzymes FatM and RAM 2 fine-tune lipid biosynthesis to promote development of arbuscular mycorrhiza". New Phytologist. 214 (4): 1631–1645. doi:10.1111/nph.14533. PMID 28380681. S2CID 3481181.
- ^ Paszkowski, Uta (October 2006). "A journey through signaling in arbuscular mycorrhizal symbioses 2006". New Phytologist. 172 (1): 35–46. doi:10.1111/j.1469-8137.2006.01840.x. PMID 16945087.
- ^ a b Bais, Harsh P.; Weir, Tiffany L.; Perry, Laura G.; Gilroy, Simon; Vivanco, Jorge M. (1 June 2006). "The Role of Root Exudates in Rhizosphere Interactions with Plants and Other Organisms". Annual Review of Plant Biology. 57 (1): 233–266. doi:10.1146/annurev.arplant.57.032905.105159. PMID 16669762.
- ^ a b c Badri, Dayakar V.; Vivanco, Jorge M. (June 2009). "Regulation and function of root exudates". Plant, Cell & Environment. 32 (6): 666–681. doi:10.1111/j.1365-3040.2009.01926.x. PMID 19143988.
- ^ a b De-la-Peña, Clelia; Badri, Dayakar V.; Loyola-Vargas, Víctor M. (2012). "Plant Root Secretions and Their Interactions with Neighbors". Secretions and Exudates in Biological Systems. Signaling and Communication in Plants. Vol. 12. pp. 1–26. doi:10.1007/978-3-642-23047-9_1. ISBN 978-3-642-23046-2.
- ^ a b c Genre, Andrea; Chabaud, Mireille; Balzergue, Coline; Puech-Pagès, Virginie; Novero, Mara; Rey, Thomas; Fournier, Joëlle; Rochange, Soizic; Bécard, Guillaume; Bonfante, Paola; Barker, David G. (April 2013). "Short-chain chitin oligomers from arbuscular mycorrhizal fungi trigger nuclear C a 2+ spiking in M edicago truncatula roots and their production is enhanced by strigolactone". New Phytologist. 198 (1): 190–202. doi:10.1111/nph.12146. hdl:2318/134858. PMID 23384011. S2CID 34009711.
- ^ a b c d Bonfante, Paola; Genre, Andrea (27 July 2010). "Mechanisms underlying beneficial plant–fungus interactions in mycorrhizal symbiosis". Nature Communications. 1 (1): 48. Bibcode:2010NatCo...1...48B. doi:10.1038/ncomms1046. hdl:2318/77771. PMID 20975705.
- ^ Maillet, Fabienne; Poinsot, Véréna; André, Olivier; Puech-Pagès, Virginie; Haouy, Alexandra; Gueunier, Monique; Cromer, Laurence; Giraudet, Delphine; Formey, Damien; Niebel, Andreas; Martinez, Eduardo Andres; Driguez, Hugues; Bécard, Guillaume; Dénarié, Jean (2011). "Fungal lipochitooligosaccharide symbiotic signals in arbuscular mycorrhiza". Nature. 469 (7328): 58–63. Bibcode:2011Natur.469...58M. doi:10.1038/nature09622. PMID 21209659. S2CID 4373531.
- ^ Sun, Jongho; Miller, J. Benjamin; Granqvist, Emma; Wiley-Kalil, Audrey; Gobbato, Enrico; Maillet, Fabienne; Cottaz, Sylvain; Samain, Eric; Venkateshwaran, Muthusubramanian; Fort, Sébastien; Morris, Richard J.; Ané, Jean-Michel; Dénarié, Jean; Oldroyd, Giles E.D. (9 April 2015). "Activation of Symbiosis Signaling by Arbuscular Mycorrhizal Fungi in Legumes and Rice". The Plant Cell. 27 (3): 823–838. doi:10.1105/tpc.114.131326. PMC 4558648. PMID 25724637.
- ^ Genre, A.; Bonfante, P. (February 2020). "A Rice Receptor for Mycorrhizal Fungal Signals Opens New Opportunities for the Development of Sustainable Agricultural Practices". Molecular Plant. 13 (2): 181–183. doi:10.1016/j.molp.2020.01.009. hdl:2318/1726475. PMID 31981734. S2CID 210912425.
- ^ a b Miyata, Kana; Kozaki, Toshinori; Kouzai, Yusuke; Ozawa, Kenjirou; Ishii, Kazuo; Asamizu, Erika; Okabe, Yoshihiro; Umehara, Yosuke; Miyamoto, Ayano; Kobae, Yoshihiro; Akiyama, Kohki; Kaku, Hanae; Nishizawa, Yoko; Shibuya, Naoto; Nakagawa, Tomomi (November 2014). "The Bifunctional Plant Receptor, OsCERK1, Regulates Both Chitin-Triggered Immunity and Arbuscular Mycorrhizal Symbiosis in Rice". Plant and Cell Physiology. 55 (11): 1864–1872. doi:10.1093/pcp/pcu129. PMID 25231970.
- ^ Miyata, Kana; Hosotani, Moe; Akamatsu, Akira; Takeda, Naoya; Jiang, Wendi; Sugiyama, Taisei; Takaoka, Ryou; Matsumoto, Kotarou; Abe, Satsuki; Shibuya, Naoto; Kaku, Hanae (2023-04-17). "OsSYMRK Plays an Essential Role in AM Symbiosis in Rice ( Oryza sativa )". Plant and Cell Physiology. 64 (4): 378–391. doi:10.1093/pcp/pcad006. ISSN 0032-0781. PMID 36688592.
- ^ Venkateshwaran, Muthusubramanian; Jayaraman, Dhileepkumar; Chabaud, Mireille; Genre, Andrea; Balloon, Allison J.; Maeda, Junko; Forshey, Kari; den Os, Désirée; Kwiecien, Nicholas W.; Coon, Joshua J.; Barker, David G.; Ané, Jean-Michel (4 August 2015). "A role for the mevalonate pathway in early plant symbiotic signaling". Proceedings of the National Academy of Sciences. 112 (31): 9781–9786. Bibcode:2015PNAS..112.9781V. doi:10.1073/pnas.1413762112. PMC 4534228. PMID 26199419.
- ^ Charpentier, Myriam; Oldroyd, Giles E.D. (8 October 2013). "Nuclear Calcium Signaling in Plants". Plant Physiology. 163 (2): 496–503. doi:10.1104/pp.113.220863. PMC 3793031. PMID 23749852.
- ^ a b Saeed, Wajeeha; Naseem, Saadia; Ali, Zahid (28 August 2017). "Strigolactones Biosynthesis and Their Role in Abiotic Stress Resilience in Plants: A Critical Review". Frontiers in Plant Science. 8: 1487. doi:10.3389/fpls.2017.01487. PMC 5581504. PMID 28894457.
- ^ a b Al-Babili, Salim; Bouwmeester, Harro J. (29 April 2015). "Strigolactones, a Novel Carotenoid-Derived Plant Hormone". Annual Review of Plant Biology. 66 (1): 161–186. doi:10.1146/annurev-arplant-043014-114759. PMID 25621512.
- ^ Gutjahr, Caroline; Gobbato, Enrico; Choi, Jeongmin; Riemann, Michael; Johnston, Matthew G.; Summers, William; Carbonnel, Samy; Mansfield, Catherine; Yang, Shu-Yi; Nadal, Marina; Acosta, Ivan; Takano, Makoto; Jiao, Wen-Biao; Schneeberger, Korbinian; Kelly, Krystyna A. (2015-12-18). "Rice perception of symbiotic arbuscular mycorrhizal fungi requires the karrikin receptor complex". Science. 350 (6267): 1521–1524. doi:10.1126/science.aac9715. ISSN 0036-8075. PMID 26680197. S2CID 206641200.
- ^ Nadal, Marina; Sawers, Ruairidh; Naseem, Shamoon; Bassin, Barbara; Kulicke, Corinna; Sharman, Abigail; An, Gynheung; An, Kyungsook; Ahern, Kevin R.; Romag, Amanda; Brutnell, Thomas P.; Gutjahr, Caroline; Geldner, Niko; Roux, Christophe; Martinoia, Enrico (2017-05-26). "An N-acetylglucosamine transporter required for arbuscular mycorrhizal symbioses in rice and maize". Nature Plants. 3 (6): 17073. doi:10.1038/nplants.2017.73. ISSN 2055-0278. PMC 5685555. PMID 28548655.
- ^ a b Nakagawa, Tomomi; Imaizumi-Anraku, Haruko (December 2015). "Rice arbuscular mycorrhiza as a tool to study the molecular mechanisms of fungal symbiosis and a potential target to increase productivity". Rice. 8 (1): 32. doi:10.1186/s12284-015-0067-0. PMC 4626465. PMID 26516078.
- ^ a b Rasmussen, S. R.; Füchtbauer, W.; Novero, M.; Volpe, V.; Malkov, N.; Genre, A.; Bonfante, P.; Stougaard, J.; Radutoiu, S. (20 July 2016). "Intraradical colonization by arbuscular mycorrhizal fungi triggers induction of a lipochitooligosaccharide receptor". Scientific Reports. 6 (1): 29733. Bibcode:2016NatSR...629733R. doi:10.1038/srep29733. PMC 4951684. PMID 27435342.
- ^ Genre, A.; Bonfante, P. (February 2020). "A Rice Receptor for Mycorrhizal Fungal Signals Opens New Opportunities for the Development of Sustainable Agricultural Practices". Molecular Plant. 13 (2): 181–183. doi:10.1016/j.molp.2020.01.009. hdl:2318/1726475. PMID 31981734. S2CID 210912425.
- ^ Miyata, Kana; Kozaki, Toshinori; Kouzai, Yusuke; Ozawa, Kenjirou; Ishii, Kazuo; Asamizu, Erika; Okabe, Yoshihiro; Umehara, Yosuke; Miyamoto, Ayano; Kobae, Yoshihiro; Akiyama, Kohki; Kaku, Hanae; Nishizawa, Yoko; Shibuya, Naoto; Nakagawa, Tomomi (November 2014). "The Bifunctional Plant Receptor, OsCERK1, Regulates Both Chitin-Triggered Immunity and Arbuscular Mycorrhizal Symbiosis in Rice". Plant and Cell Physiology. 55 (11): 1864–1872. doi:10.1093/pcp/pcu129. ISSN 1471-9053. PMID 25231970.
- ^ a b c d Buendia, Luis; Girardin, Ariane; Wang, Tongming; Cottret, Ludovic; Lefebvre, Benoit (2018-10-24). "LysM Receptor-Like Kinase and LysM Receptor-Like Protein Families: An Update on Phylogeny and Functional Characterization". Frontiers in Plant Science. 9: 1531. doi:10.3389/fpls.2018.01531. ISSN 1664-462X. PMC 6207691. PMID 30405668.
- ^ He, Jiangman; Zhang, Chi; Dai, Huiling; Liu, Huan; Zhang, Xiaowei; Yang, Jun; Chen, Xi; Zhu, Yayun; Wang, Dapeng; Qi, Xiaofeng; Li, Weichao; Wang, Zhihui; An, Guoyong; Yu, Nan; He, Zuhua (December 2019). "A LysM Receptor Heteromer Mediates Perception of Arbuscular Mycorrhizal Symbiotic Signal in Rice". Molecular Plant. 12 (12): 1561–1576. doi:10.1016/j.molp.2019.10.015. PMID 31706032. S2CID 207966375.
- ^ a b c d Nakagawa, Tomomi; Imaizumi-Anraku, Haruko (December 2015). "Rice arbuscular mycorrhiza as a tool to study the molecular mechanisms of fungal symbiosis and a potential target to increase productivity". Rice. 8 (1): 32. doi:10.1186/s12284-015-0067-0. ISSN 1939-8425. PMC 4626465. PMID 26516078.
- ^ Paszkowski, Uta (October 2006). "A journey through signaling in arbuscular mycorrhizal symbioses 2006". New Phytologist. 172 (1): 35–46. doi:10.1111/j.1469-8137.2006.01840.x. ISSN 0028-646X. PMID 16945087.
- ^ Kosuta, Sonja; Chabaud, Mireille; Lougnon, Géraldine; Gough, Clare; Dénarié, Jean; Barker, David G.; Bécard, Guillaume (2003-03-01). "A Diffusible Factor from Arbuscular Mycorrhizal Fungi Induces Symbiosis-Specific MtENOD11 Expression in Roots of Medicago truncatula". Plant Physiology. 131 (3): 952–962. doi:10.1104/pp.011882. ISSN 1532-2548. PMC 166861. PMID 12644648.
- ^ MacLean, Allyson M.; Bravo, Armando; Harrison, Maria J. (October 2017). "Plant Signaling and Metabolic Pathways Enabling Arbuscular Mycorrhizal Symbiosis". The Plant Cell. 29 (10): 2319–2335. doi:10.1105/tpc.17.00555. ISSN 1040-4651. PMC 5940448. PMID 28855333.
- ^ Genre, A.; Bonfante, P. (February 2020). "A Rice Receptor for Mycorrhizal Fungal Signals Opens New Opportunities for the Development of Sustainable Agricultural Practices". Molecular Plant. 13 (2): 181–183. doi:10.1016/j.molp.2020.01.009. hdl:2318/1726475. PMID 31981734. S2CID 210912425.
- ^ Miyata, Kana; Kozaki, Toshinori; Kouzai, Yusuke; Ozawa, Kenjirou; Ishii, Kazuo; Asamizu, Erika; Okabe, Yoshihiro; Umehara, Yosuke; Miyamoto, Ayano; Kobae, Yoshihiro; Akiyama, Kohki; Kaku, Hanae; Nishizawa, Yoko; Shibuya, Naoto; Nakagawa, Tomomi (November 2014). "The Bifunctional Plant Receptor, OsCERK1, Regulates Both Chitin-Triggered Immunity and Arbuscular Mycorrhizal Symbiosis in Rice". Plant and Cell Physiology. 55 (11): 1864–1872. doi:10.1093/pcp/pcu129. ISSN 1471-9053. PMID 25231970.
- ^ 3. MacLean AM, Bravo A, Harrison MJ. Plant signaling and metabolic pathways enabling arbuscularmycorrhizal symbiosis. The Plant Cell. 2017; 29(10):2319-2335. https://doi.org/10.1105/tpc.17.00555
- ^ a b c d e Bonfante, Paola; Genre, Andrea (2010-07-27). "Mechanisms underlying beneficial plant–fungus interactions in mycorrhizal symbiosis". Nature Communications. 1 (1): 48. Bibcode:2010NatCo...1...48B. doi:10.1038/ncomms1046. hdl:2318/77771. ISSN 2041-1723. PMID 20975705.
- ^ a b c d e f g h i j k l m n o MacLean, Allyson M.; Bravo, Armando; Harrison, Maria J. (October 2017). "Plant Signaling and Metabolic Pathways Enabling Arbuscular Mycorrhizal Symbiosis". The Plant Cell. 29 (10): 2319–2335. doi:10.1105/tpc.17.00555. ISSN 1040-4651. PMC 5940448. PMID 28855333.
- ^ a b Venkateshwaran, Muthusubramanian; Jayaraman, Dhileepkumar; Chabaud, Mireille; Genre, Andrea; Balloon, Allison J.; Maeda, Junko; Forshey, Kari; den Os, Désirée; Kwiecien, Nicholas W.; Coon, Joshua J.; Barker, David G.; Ané, Jean-Michel (2015-08-04). "A role for the mevalonate pathway in early plant symbiotic signaling". Proceedings of the National Academy of Sciences. 112 (31): 9781–9786. Bibcode:2015PNAS..112.9781V. doi:10.1073/pnas.1413762112. ISSN 0027-8424. PMC 4534228. PMID 26199419.
- ^ Kosuta, Sonja; Chabaud, Mireille; Lougnon, Géraldine; Gough, Clare; Dénarié, Jean; Barker, David G.; Bécard, Guillaume (2003-03-01). "A Diffusible Factor from Arbuscular Mycorrhizal Fungi Induces Symbiosis-Specific MtENOD11 Expression in Roots of Medicago truncatula". Plant Physiology. 131 (3): 952–962. doi:10.1104/pp.011882. ISSN 1532-2548. PMC 166861. PMID 12644648.
- ^ Pimprikar, Priya; Gutjahr, Caroline (2018-04-01). "Transcriptional Regulation of Arbuscular Mycorrhiza Development". Plant and Cell Physiology. 59 (4): 678–695. doi:10.1093/pcp/pcy024. ISSN 0032-0781. PMID 29425360.
- ^ Hirsch, Sibylle; Oldroyd, Giles E.D. (August 2009). "GRAS-domain transcription factors that regulate plant development". Plant Signaling & Behavior. 4 (8): 698–700. doi:10.4161/psb.4.8.9176. ISSN 1559-2324. PMC 2801379. PMID 19820314.
- ^ Ho-Plágaro, Tania; García-Garrido, José Manuel (2022). "Multifarious and Interactive Roles of GRAS Transcription Factors During Arbuscular Mycorrhiza Development". Frontiers in Plant Science. 13. doi:10.3389/fpls.2022.836213. ISSN 1664-462X. PMC 8996055. PMID 35419017.
- ^ Mohanta, Tapan Kumar; Bae, Hanhong (January 2015). "Functional genomics and signaling events in mycorrhizal symbiosis". Journal of Plant Interactions. 10 (1): 21–40. doi:10.1080/17429145.2015.1005180. ISSN 1742-9145. S2CID 85269506.
- ^ Paszkowski, Uta (October 2006). "A journey through signaling in arbuscular mycorrhizal symbioses 2006". New Phytologist. 172 (1): 35–46. doi:10.1111/j.1469-8137.2006.01840.x. ISSN 0028-646X. PMID 16945087.
- ^ a b Luginbuehl, Leonie H.; Menard, Guillaume N.; Kurup, Smita; Van Erp, Harrie; Radhakrishnan, Guru V.; Breakspear, Andrew; Oldroyd, Giles E. D.; Eastmond, Peter J. (2017-06-16). "Fatty acids in arbuscular mycorrhizal fungi are synthesized by the host plant". Science. 356 (6343): 1175–1178. doi:10.1126/science.aan0081. ISSN 0036-8075. PMID 28596311. S2CID 206658467.
- ^ Bravo, Armando; Brands, Mathias; Wewer, Vera; Dörmann, Peter; Harrison, Maria J. (June 2017). "Arbuscular mycorrhiza-specific enzymes FatM and RAM 2 fine-tune lipid biosynthesis to promote development of arbuscular mycorrhiza". New Phytologist. 214 (4): 1631–1645. doi:10.1111/nph.14533. ISSN 0028-646X. PMID 28380681. S2CID 3481181.
- ^ Guo, Rui; Wu, Ya-Nan; Liu, Cheng-Chen; Liu, Ying-Na; Tian, Li; Cheng, Jian-Fei; Pan, Zhiyong; Wang, Dong; Wang, Bin (2022). "OsADK1, a novel kinase regulating arbuscular mycorrhizal symbiosis in rice". New Phytologist. 234 (1): 256–268. doi:10.1111/nph.17979. ISSN 0028-646X. PMID 35133010.
- ^ Leng, Junchen; Wei, Xiaotong; Jin, Xinyi; Wang, Longxiang; Fan, Kai; Zou, Ke; Zheng, Zichao; Saridis, Georgios; Zhao, Ningkang; Zhou, Dan; Duanmu, Deqiang; Wang, Ertao; Cui, Haitao; Bucher, Marcel; Xue, Li (2023). "Arbuscular Mycorrhiza-Induced Kinases AMK8 and AMK24 associate with the receptor-like kinase KINASE3 to regulate arbuscular mycorrhizal symbiosis in Lotus japonicus". The Plant Cell. pp. 2006–2026. doi:10.1093/plcell/koad050. PMC 10226568. PMID 36808553. Retrieved 2023-10-30.
- ^ Roth, Ronelle; Chiapello, Marco; Montero, Héctor; Gehrig, Peter; Grossmann, Jonas; O’Holleran, Kevin; Hartken, Denise; Walters, Fergus; Yang, Shu-Yi; Hillmer, Stefan; Schumacher, Karin; Bowden, Sarah; Craze, Melanie; Wallington, Emma J.; Miyao, Akio (2018-11-08). "A rice Serine/Threonine receptor-like kinase regulates arbuscular mycorrhizal symbiosis at the peri-arbuscular membrane". Nature Communications. 9 (1): 4677. doi:10.1038/s41467-018-06865-z. ISSN 2041-1723. PMC 6224560. PMID 30410018.
- ^ Montero, Héctor; Lee, Tak; Pucker, Boas; Ferreras-Garrucho, Gabriel; Oldroyd, Giles; Brockington, Samuel F.; Miyao, Akio; Paszkowski, Uta (2021-06-22). "A mycorrhiza-associated receptor-like kinase with an ancient origin in the green lineage". Proceedings of the National Academy of Sciences. 118 (25). doi:10.1073/pnas.2105281118. ISSN 0027-8424. PMC 8237591. PMID 34161289.
- ^ Wang, Wanxiao; Shi, Jincai; Xie, Qiujin; Jiang, Yina; Yu, Nan; Wang, Ertao (September 2017). "Nutrient Exchange and Regulation in Arbuscular Mycorrhizal Symbiosis". Molecular Plant. 10 (9): 1147–1158. doi:10.1016/j.molp.2017.07.012. PMID 28782719.
- ^ Barman, Jagnaseni; Samanta, Aveek; Saha, Babita; Datta, Siraj (December 2016). "Mycorrhiza: The oldest association between plant and fungi". Resonance. 21 (12): 1093–1104. doi:10.1007/s12045-016-0421-6. ISSN 0971-8044. S2CID 109932925.
- ^ Ercolin, Flavia; Reinhardt, Didier (July 2011). "Successful joint ventures of plants: arbuscular mycorrhiza and beyond" (PDF). Trends in Plant Science. 16 (7): 356–362. doi:10.1016/j.tplants.2011.03.006. PMID 21459657.
- ^ De-la-Peña, Clelia; Badri, Dayakar V.; Loyola-Vargas, Víctor M. (2012), Vivanco, Jorge M.; Baluška, František (eds.), "Plant Root Secretions and Their Interactions with Neighbors", Secretions and Exudates in Biological Systems, Signaling and Communication in Plants, vol. 12, Berlin, Heidelberg: Springer Berlin Heidelberg, pp. 1–26, doi:10.1007/978-3-642-23047-9_1, ISBN 978-3-642-23046-2, retrieved 2023-10-24