An arbuscular mycorrhiza (AM) (plural mycorrhizae) is a type of mycorrhiza in which the symbiont fungus (AM fungi, or AMF) penetrates the cortical cells of the roots of a vascular plant forming arbuscules. Arbuscular mycorrhiza is a type of endomycorrhiza along with ericoid mycorrhiza and orchid mycorrhiza (not to be confused with ectomycorrhiza). They are characterized by the formation of unique tree-like structures, the arbuscules.[1] In addition, globular storage structures called vesicles are often encountered.



Arbuscular mycorrhizae are formed by fungi in the subphylum Glomeromycotina. This subphylum, along with the Mortierellomycotina, and Mucoromycotina, form the phylum Mucoromycota, a sister clade of the more well-known and diverse dikaryan fungi.[2]
AM fungi help plants to capture nutrients such as phosphorus, sulfur, nitrogen and micronutrients from the soil. It is believed that the development of the arbuscular mycorrhizal symbiosis played a crucial role in the initial colonisation of land by plants and in the evolution of the vascular plants.[3] It has been said that it is quicker to list the plants that do not form endomycorrhizae than those that do.[4] This symbiosis is a highly evolved mutualistic relationship found between fungi and plants, the most prevalent plant symbiosis known,[5] and AMF is found in 80% of vascular plant families in existence today.[6]
Previously this type of mycorrhizal associations were called 'Vesicular arbuscular mycorrhiza (VAM)', but since some members of these fungi do not produce any vesicles, such as the members of Gigasporaceae; the term has been changed to 'Arbuscular Mycorrhizae' to include them.[7][8]
Advances in research on mycorrhizal physiology and ecology since the 1970s have led to a greater understanding of the multiple roles of AMF in the ecosystem. An example is the important contribution of the glue-like protein glomalin to soil structure (see below). This knowledge is applicable to human endeavors of ecosystem management, ecosystem restoration, and agriculture.
Evolution of mycorrhizal symbiosis
editPaleobiology
editBoth paleobiological and molecular evidence indicate that AM is an ancient symbiosis that originated at least 460 million years ago. AM symbiosis is ubiquitous among land plants, which suggests that mycorrhizas were present in the early ancestors of extant land plants. This positive association with plants may have facilitated the development of land plants.[5]
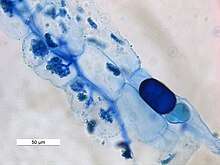
The Rhynie chert of the lower Devonian has yielded fossils of the earliest land plants in which AM fungi have been observed.[10] The fossilized plants containing mycorrhizal fungi were preserved in silica.
The Early Devonian saw the development of terrestrial flora. Plants of the Rhynie chert from the Lower Devonian (400 m.yrs ago) were found to contain structures resembling vesicles and spores of present Glomus species. Colonized fossil roots have been observed in Aglaophyton major and Rhynia, which are ancient plants possessing characteristics of vascular plants and bryophytes with primitive protostelic rhizomes.[10]
Intraradical mycelium was observed in root intracellular spaces, and arbuscules were observed in the layer thin wall cells similar to palisade parenchyma. The fossil arbuscules appear very similar to those of existing AMF.[10] The cells containing arbuscules have thickened walls, which are also observed in extant colonized cells.
Mycorrhizas from the Miocene exhibit a vesicular morphology closely resembling that of present Glomerales. This conserved morphology may reflect the ready availability of nutrients provided by the plant hosts in both modern and Miocene mutualisms.[11] However, it can be argued that the efficacy of signaling processes is likely to have evolved since the Miocene, and this can not be detected in the fossil record. A finetuning of the signaling processes would improve coordination and nutrient exchange between symbionts while increasing the fitness of both the fungi and the plant symbionts.
The nature of the relationship between plants and the ancestors of arbuscular mycorrhizal fungi is contentious. Two hypotheses are:
- Mycorrhizal symbiosis evolved from a parasitic interaction that developed into a mutually beneficial relationship.
- Mycorrhizal fungi developed from saprobic fungi that became endosymbiotic.[10]
Both saprotrophs and biotrophs were found in the Rhynie Chert, but there is little evidence to support either hypothesis.
There is some fossil evidence that suggests that the parasitic fungi did not kill the host cells immediately upon invasion, although a response to the invasion was observed in the host cells. This response may have evolved into the chemical signaling processes required for symbiosis.[10]
In both cases, the symbiotic plant-fungi interaction is thought to have evolved from a relationship in which the fungi was taking nutrients from the plant into a symbiotic relationship where the plant and fungi exchange nutrients.
The ancient plants did not have true roots. Strullu-Derrien and Strullu proposed the term 'Paramycorrhizae' for the mycorrhizae that infected the rhizome or shoot or thalli, and 'Eumycorrhizae' that infects true roots.[12][13][14] These structures were reported in both sporophytes and gametophytes of the early land plants.[13]
Molecular evidence
editIncreased interest in mycorrhizal symbiosis and the development of sophisticated molecular techniques has led to the rapid development of genetic evidence. Wang et al. (2010) investigated plant genes involved in communication with order Glomales fungal partners (DMI1, DMI3, IPD3).[15][16] These three genes could be sequenced from all major clades of modern land plants, including liverworts, the most basal group, and phylogeny of the three genes proved to agree with then current land plant phylogenies. This implies that mycorrhizal genes must have been present in the common ancestor of land plants, and that they must have been vertically inherited since plants colonized land.[15]
AM fungi and cyanobacteria symbiosis
editIt was revealed that AM fungi have the bacterial type core enzyme (ribonuclease III) of the sRNA processing mechanism, probably by the process of horizontal gene transfer from a cyanobacterial ancestor, and possibly related to symbiosis,.[17] This finding of a genetic fossil inside AM fungi raises the possibility of an intimate relationship between AM fungi and cyanobacterial ancestors. A similar Geosiphon–Nostoc symbiosis was previously reported.[18]
Circadian clock evolution in AM fungi
editSurprisingly, despite their long evolution as an underground partner of plant roots, whose environment is far from light or temperature fluctuation, AMF still have a conserved circadian clock whose fungal circadian oscillator (frq) is activated by the blue light, similar to the model circadian fungus Neurospora crassa.[19] The proven conservation of a circadian clock and output genes in R. irregulare opens the door to the study of circadian clocks in the fungal partner of AM symbiosis. The same research characterized the AMF frq gene,[19] which is the first frq gene identified outgroup of Dikarya, and suggests the frq gene evolution in the fungal kingdom is much older than previously thought.
Physiology
editPresymbiosis
editThe development of the AM fungi prior to root colonization, known as presymbiosis, consists of three stages: spore germination, hyphal growth, host recognition and appressorium formation.
Spore germination
editSpores of the AM fungi are thick-walled multi-nucleate resting structures.[21] The germination of the spore does not depend on the plant, as spores have been germinated under experimental conditions in the absence of plants both in vitro and in soil. However, the rate of germination can be increased by host root exudates.[22] AM fungal spores germinate given suitable conditions of the soil matrix, temperature, carbon dioxide concentration, pH, and phosphorus concentration.[21]
Hyphal growth
editThe growth of AM hyphae through the soil is controlled by host root exudates known as strigolactones, and the soil phosphorus concentration.[23] Low-phosphorus concentrations in the soil increase hyphal growth and branching as well as induce plant exudation of compounds that control hyphal branching intensity.[22][24]
The branching of AM fungal hyphae grown in phosphorus media of 1 mM is significantly reduced, but the length of the germ tube and total hyphal growth were not affected. A concentration of 10 mM phosphorus inhibited both hyphal growth and branching. This phosphorus concentration occurs in natural soil conditions and could thus contribute to reduced mycorrhizal colonization.[24]
Host recognition
editRoot exudates from AMF host plants grown in a liquid medium with and without phosphorus have been shown to affect hyphal growth. Spores of Gigaspora margarita were grown in host plant exudates. Hyphae of fungi grown in the exudates from roots starved of phosphorus grew more and produced tertiary branches compared to those grown in exudates from plants given adequate phosphorus. When the growth-promoting root exudates were added in low concentration, the AM fungi produced scattered long branches. As the concentration of exudates was increased, the fungi produced more tightly clustered branches. At the highest-concentration arbuscules, the AMF structures of phosphorus exchange were formed.[24]
This chemotaxic fungal response to the host plants exudates is thought to increase the efficacy of host root colonization in low-phosphorus soils.[22] It is an adaptation for fungi to efficiently explore the soil in search of a suitable plant host.[24]
Further evidence that arbuscular mycorrhizal fungi exhibit host-specific chemotaxis, that enable hyphal growth toward the roots of a potential host plant: Spores of Glomus mosseae were separated from the roots of a host plant, nonhost plants, and dead host plant by a membrane permeable only to hyphae. In the treatment with the host plant, the fungi crossed the membrane and always emerged within 800 μm of the root, but not in the treatments with nonhost plants and dead plants.[25]
Molecular techniques have been used to understand the signaling pathways between arbuscular mycorrhizae and plant roots. In 2003 it was shown how the AM undergoes physiological changes in the presence of exudates from potential host plant roots, to colonize it. Host plant root exudates trigger and turn on AM fungal genes required for the respiration of spore carbon compounds. In experiments, transcription rate of 10 genes increased half-hour after exposure and at an even greater rate after 1 hour. after 4 hours exposure AM respond with morphological growth. Genes isolated from that time are involved in mitochondrial activity and enzyme production. The fungal respiration rate, measured by O2 consumption rate, increased by 30% 3 hours after exposure to root exudates, indicating that host plant root exudates stimulate AMF spore mitochondrial activity. It may be part of a fungal regulatory mechanism that conserves spore energy for efficient growth and the hyphal branching upon receiving signals from a potential host plant.[26]
Appressorium
editWhen arbuscular mycorrhizal fungal hyphae encounter the root of a host plant, an appressorium or 'infection structure' forms on the root epidermis. From this structure hyphae can penetrate into the host's parenchyma cortex.[27] AM need no chemical signals from the plant to form the appressoria. AM fungi could form appressoria on the cell walls of "ghost" cells in which the protoplast had been removed to eliminate signaling between the fungi and the plant host. However, the hyphae did not further penetrate the cells and grow in toward the root cortex, which indicates that signaling between symbionts is required for further growth once appressoria are formed.[22]
Symbiosis
editOnce inside the parenchyma, the fungus forms highly branched structures for nutrient exchange with the plant called arbuscules.[27] These are the distinguishing structures of arbuscular mycorrhizal fungus. Arbuscules are the sites of exchange for phosphorus, carbon, water, and other nutrients.[21] There are two forms: Paris type is characterized by the growth of hyphae from one cell to the next; and Arum type is characterized by the growth of hyphae in the space between plant cells.[28] The choice between Paris type and Arum type is primarily determined by the host plant family, although some families or species are capable of either type.[29][30]
The host plant exerts a control over the intercellular hyphal proliferation and arbuscule formation. There is a decondensation of the plant's chromatin, which indicates increased transcription of the plant's DNA in arbuscule-containing cells.[27] Major modifications are required in the plant host cell to accommodate the arbuscules. The vacuoles shrink and other cellular organelles proliferate. The plant cell cytoskeleton is reorganized around the arbuscules.
There are two other types of hyphae that originate from the colonized host plant root. Once colonization has occurred, short-lived runner hyphae grow from the plant root into the soil. These are the hyphae that take up phosphorus and micronutrients, which are conferred to the plant. AM fungal hyphae have a high surface-to-volume ratio, making their absorptive ability greater than that of plant roots.[31] AMF hyphae are also finer than roots and can enter into pores of the soil that are inaccessible to roots.[32] The fourth type of AMF hyphae grows from the roots and colonizes other host plant roots. The four types of hyphae are morphologically distinct.[21]
Nutrient uptake and exchange
editAM fungi are obligate symbionts. They have limited saprobic ability and depend on the plant for their carbon nutrition.[33] AM fungi take up the products of the plant host's photosynthesis as hexoses.
Carbon transfer from plant to fungi may occur through the arbuscules or intraradical hyphae.[34] Secondary synthesis from the hexoses by AM occurs in the intraradical mycelium. Inside the mycelium, hexose is converted to trehalose and glycogen. Trehalose and glycogen are carbon storage forms that can be rapidly synthesized and degraded and may buffer the intracellular sugar concentrations.[34] The intraradical hexose enters the oxidative pentose phosphate pathway, which produces pentose for nucleic acids.
Lipid biosynthesis also occurs in the intraradical mycelium. Lipids are then stored or exported to extraradical hyphae where they may be stored or metabolized. The breakdown of lipids into hexoses, known as gluconeogenesis, occurs in the extraradical mycelium.[34] Approximately 25% of the carbon translocated from the plant to the fungi is stored in the extraradical hyphae.[35] Up to 20% of the host plant's carbon may be transferred to the AM fungi.[34] This represents the host plant's considerable carbon investment in mycorrhizal network and contribution to the below-ground organic carbon pool.
Increasing the plant's carbon supply to the AM fungi increases uptake and transfer of phosphorus from fungi to plant.[36] Likewise, phosphorus uptake and transfer is lowered when the photosynthate supplied to the fungi is decreased. Species of AMF differ in their abilities to supply the plant with phosphorus.[37] In some cases, arbuscular mycorrhizae are poor symbionts, providing little phosphorus while taking relatively high amounts of carbon.[37]
The main benefit of mycorrhizas to plants has been attributed to increased uptake of nutrients, especially phosphorus. This may be due to increased surface area in contact with soil, increased movement of nutrients into mycorrhizae, a modified root environment, and increased storage.[32] Mycorrhizas can be much more efficient than plant roots at taking up phosphorus. Phosphorus travels to the root or via diffusion and hyphae reduce the distance required for diffusion, thus increasing uptake. The rate of phosphorus flowing into mycorrhizae can be up to six times that of the root hairs.[32] In some cases, the role of phosphorus uptake can be completely taken over by the mycorrhizal network, and all of the plant's phosphorus may be of hyphal origin.[37] Less is known about the role of nitrogen nutrition in the arbuscular mycorrhizal system and its impact on the symbiosis and community. While significant advances have been made in elucidating the mechanisms of this complex interaction, much investigation remains to be done.
Mycorrhizal activity increases the phosphorus concentration available in the rhizosphere. Mycorrhizae lower the root zone pH by selective uptake of NH4+ (ammonium-ions) and by releasing H+ ions. Decreased soil pH increases the solubility of phosphorus precipitates. The hyphal NH4+ uptake also increases the nitrogen flow to the plant as the soil's inner surfaces absorb ammonium and distribute it by diffusion.[35]
Meiosis and recombination
editAM fungi have been regarded as asexual because they lack observable sexual structures.[38] However, homologs of 51 genes that function in meiosis, including seven meiosis-specific genes were found to be conserved in the genomes of several AMF species, suggesting that these supposedly ancient asexual fungi may be capable of undergoing conventional meiosis.[39] Furthermore, in Rhizophagus irregularis genetic exchange involving reciprocal recombination was found to occur in dikaryons between haploid genomes.[38]
Mechanism of colonization
editRecent research has shown that AM fungi release a diffusional factor, known as the myc factor, which activates the nodulation factor's inducible gene MtEnod11. This is the same gene involved in establishing symbiosis with the nitrogen fixing, rhizobial bacteria (Kosuta et al. 2003). The factor was first identified by Fabienne Maillet and coworkers[40] in a groundbreaking work published in Nature, where they have extracted three hundred litre mycorrhized carrot roots and exudates from 40 million germinating spores of Rhizophagus irregularis and purified the active fraction. They demonstrated this active principle is lipo-chito-oligosaccharide in nature.
Recognition of Myc factors triggers the common symbiotic signaling pathway (CSSP) that eventually leads to plant's accommodation programme to provide hostage to the arbuscular mycorrhizae.
The Common Symbiotic Signaling Pathway (CSSP) is a Signaling cascade in plants that seen to be activated in both NOD-factor perception (for nodule forming Rhizobia), as well as found in MYC-factor perception that are released from Arbuscular mycorrhizal fungi. The pathway is distinguished from the pathogen recognition pathways, but may have some common receptors involved in both pathogen recognition as well as CSSP. A recent work[41] by Kevin Cope and colleagues shown that possibly other type of mycorrhizae may involve the CSSP components such as Myc-factor recognition.
The AMF colonization requires the following chain[42] of events that can be roughly divided into following steps -
- 1.The Pre-Contact Signaling,
- 2. The CSSP
- 2.A. Perception
- 2.B. Transmission
- 2.C. Transcription
- 3. The Accommodation program
Ecology
editBiogeography
editArbuscular mycorrhizal fungi are most frequent in plants growing on mineral soils, and are of extreme importance for plants growing in nutrient-deficient substrates such as in volcanic soil and sand dune environments. The populations of AM fungi is greatest in plant communities with high diversity such as tropical rainforests and temperate grasslands where they have many potential host plants and can take advantage of their ability to colonize a broad host range.[43] There is a lower incidence of mycorrhizal colonization in very arid or nutrient-rich soils. Mycorrhizas have been observed in aquatic habitats; however, waterlogged soils have been shown to decrease colonization in some species.[43] Arbuscular mycorrhizal fungi are found in 80% of plant species[44] and have been surveyed on all continents except Antarctica.[45][46] The biogeography of glomeromycota is influenced by dispersal limitation,[47] environmental factors such as climate,[45] soil series and soil pH,[46] soil nutrients[48] and plant community.[45][49] While evidence from 2000 suggests that AM fungi are not specialists on their host species,[50] studies as of 2002 have indicated that at least some fungi taxa are host specialists.[51] The ecology of Mucoromycotinian fungi, which form "fine root endophyte" arbuscular mycorrhizas is largely unknown.
Response to plant communities
editThe specificity, host range, and degree of colonization of mycorrhizal fungi are difficult to analyze in the field due to the complexity of interactions between the fungi within a root and within the system. There is no clear evidence to suggest that arbuscular mycorrhizal fungi exhibit specificity for colonization of potential AM host plant species as do fungal pathogens for their host plants.[43] This may be due to the opposite selective pressure involved.
In pathogenic relations, the host plant benefits from mutations that prevent colonization, whereas, in a mutualistic symbiotic relationship, the plant benefits from mutation that allow for colonization by AMF.[43] However, plant species differ in the extent and dependence on colonization by certain AM fungi, and some plants may be facultative mycotrophs, while others may be obligate mycotrophs.[43] Recently, mycorrhizal status has been linked to plant distributions,[52] with obligate mycorrhizal plants occupying warmer, drier habitats while facultative mycorrhizal plants occupy larger ranges of habitats.
The ability of the same AM fungi to colonize many species of plants has ecological implications. Plants of different species can be linked underground to a common mycelial network.[43] One plant may provide the photosynthate carbon for the establishment of the mycelial network that another plant of a different species can utilize for mineral uptake. This implies that arbuscular mycorrhizae are able to balance below-ground intra–and interspecific plant interactions.[43]
Since Glomeromycota fungi live inside plant roots, they can be influenced substantially by their plant host and in return affect plant communities as well. Plants can allocate up to 30% of their photosynthate carbon to AM fungi[53] and in return AM fungi can acquire up to 80% of plant phosphorus and nitrogen.[44] The diversity of AM fungal communities has been positively linked to plant diversity,[54] plant productivity[55] and herbivory.[56] Arbuscular mycorrhizal fungi can be influenced by small scale interactions with the local plant community. For example, the plant neighborhood around a focal plant can alter AM fungal communities[57] as can the order of plant establishment within sites.[58]
AM fungi and plant invasion
editDuring invasions of plant species, the AM fungal community and biomass can be drastically altered. In the majority of cases AM fungal biomass and diversity decrease with invasions.[59][60][61] However, some mycotrophic plant species may actually increase AM fungal diversity during invasion.[62]
The mycorrhizal status of invasive plant species often varies between regions. For example, in the United Kingdom and central Europe recently invasive plants are more frequently obligately mycorrhizal than expected,[52][63] while invasive plants in California were found to be less frequently mycorrhizal than expected.[64]
Interactions between AM fungi and other plant symbionts
editAll symbionts within a plant host interact, often in unpredictable ways. A 2010 meta-analysis indicated that plants colonized by both AM fungi and vertically transmitted endophytes often are larger than plants independently colonized by these symbionts.[65] However, this relationship is context-dependent as AM fungi can interact synergistically with fungal endophytes inhabiting the leaves of their host plant,[66][67] or antagonistically.[68][69][70] Similar ranges of interactions can occur between AM fungi and ectomycorrhizal fungi and dark septate endophytes.[71]
Response to environmental gradients
editArbuscular mycorrhizal fungi vary across many environmental gradients. Their tolerance to freezing and drying is known to shift between AM fungal taxa.[72] AM fungi become less prevalent and diverse at higher soil nutrient and moisture concentrations,[73] presumably because both plants allocate less carbon to AM fungi and AM fungi reallocate their resources to intraradical hyphae in these environmental conditions.[74] Over the long term, these environmental conditions can even create local adaptation between plant hosts, AM fungi and the local soil nutrient concentrations.[75] AM composition often becomes less diverse on mountain tops than at lower elevations, which is driven by the composition of plant species.[76]
AM fungi have been shown to improve plant tolerance to abiotic environmental factors such as salinity. They alleviate salt stress and benefit plant growth and productivity.[77]
Rhizosphere ecology
editThe rhizosphere is the soil zone in the immediate vicinity of a root system.
Arbuscular mycorrhizal symbiosis affects the community and diversity of other organisms in the soil. This can be directly seen by the release of exudates, or indirectly by a change in the plant species and plant exudates type and amount.[78]
Mycorrhizae diversity has been shown to increase plant species diversity as the potential number of associations increases. Dominant arbuscular mycorrhizal fungi can prevent the invasion of non-mycorrhizal plants on land where they have established symbiosis and promote their mycorrhizal host.[79]
When rhizobium bacteria are present in the soil, mycorrhizal colonization is increased due to an increase in the concentration of chemical signals involved in the establishment of symbiosis (Xie et al. 2003). Molecules similar to Nod factors were isolated from AM fungi and were shown to induce MtEnod11, lateral root formation and enhance mycorrhization.[80] Effective mycorrhizal colonization can also increase the nodulations and symbiotic nitrogen fixation in mycorrhizal legumes.[35]
The extent of arbuscular mycorrhizal colonization and species affects the bacterial population in the rhizosphere.[81] Bacterial species differ in their abilities to compete for carbon compound root exudates. A change in the amount or composition of root exudates and fungal exudates due to the existing AM mycorrhizal colonization determines the diversity and abundance of the bacterial community in the rhizosphere.[78]
The influence of AM fungi on plant root and shoot growth may also have indirect effect on the rhizosphere bacteria. AMF contributes a substantial amount of carbon to the rhizosphere through the growth and degeneration of the hyphal network. There is also evidence to suggest that AM fungi may play an important role on mediating the plant species' specific effect on the bacterial composition of the rhizosphere.[78]
Glomeromycota and global climate change
editGlobal climate change is affecting AM fungal communities and interactions between AM fungi and their plant hosts. While it is generally accepted that interactions between organisms will affect their response to global climate change, we still lack the ability to predict the outcome of these interactions in future climates.[82] In recent meta-analyses, AM fungi were found to increase plant biomass under drought conditions and decrease plant biomass under simulated nitrogen deposition studies.[83][84] Arbuscular mycorrhizal fungi themselves have been shown to increase their biomass in response to elevated atmospheric CO2.[85]
Plants lacking arbuscular mycorrhizae
editMembers of the mustard family (Brassicaceae), such as cabbage, cauliflower, canola, and crambe, do not establish arbuscular mycorrhizal fungi on their roots.[86]
Molecular genetic analyses of arbuscular mycorrhizal fungi
editIn the past ten years there have been spectacular advances in molecular genetic technologies and tools. These advances allow microbial and mycorrhizal ecologists to ask new and exciting questions about the ecological and evolutionary roles of arbuscular mycorrhizal (AM) fungi as individuals, in communities and ecosystems. Genetic analyses of AM fungi have been used to explore the genetic structure of single spores using multilocus genotyping,[87] AM fungal diversity and adaptation across multiple grassland communities,[88] all the way up to a global investigation of AM fungal diversity, which greatly increased the described molecular diversity within the phylum Glomeromycota.[89]
All the recent advances in molecular genetics clearly permit the analysis of microbial communities at much finer and functional scales and potentially with more confidence than previous methods. The classical AM fungal identification method of spore extraction from soil and further spore morphological analysis[90] is fraught with complicating issues due to the various strategies and forms of AM fungi, e.g., lack of sporulation in certain species, seasonality, high unculturability, possible misidentification (human error), and new evidence of multi-nucleate spores[91] and high genetic variation within clonal AM species.[92] Because of these various problems, in the past researchers likely misrepresented the true composition of AM fungal communities present at any one point in time or place. Additionally, by following the traditional extraction, culture and microscopic identification methods, there is no way to determine active/functioning AM fungal populations, which are likely the most important when attempting to relate plant-AM symbiotic interactions and mechanisms to ecological or ecosystem function. This is especially true in the case of root colonization analyses, which can determine percentage of roots colonized by AM fungi. The major problem with this analysis is in field soils, which contain multiple species of AM fungi in association with a target plant at the same time (see Ecology of AM). The identification of the associated fungal symbionts is impossible without the use of molecular methods. Though genetic analysis of AM fungal communities has advanced a great deal in the past decade, the methodology is not yet completely refined. Below is an overview of the methods used in molecular genetic analyses of AM fungi, along with applications to research, future directions and some of their problems.
Overview of methods
editDNA/RNA
editGenetic analyses of AM fungi from soil and root samples range in their applicability to answer ecological or phylogenetic questions. DNA analyses utilize various nuclear markers to describe AM fungi and represent different regions of the nuclear ribosomal operon (18S rRNA) found in all eukaryotic organisms. The DNA analysis of AM fungi using these markers began in the early 1990s[93] and are continuing to be developed today. The small subunit (SSU) rRNA gene, the internal transcribed spacer (ITS) gene, and the large subunit (LSU) rRNA gene are currently the most common DNA markers used. The SSU region has been used most frequently in ecological studies,[94] while the ITS and LSU regions have been predominantly used in taxonomic constructions of the phylum Glomeromycota.[95]
qPCR and qRT-PCR
editReal-time PCR or quantitative PCR (qPCR), is becoming a well-established method to quickly amplify and simultaneously quantify targeted AM fungal DNA from biological samples (plant roots or soils). Fairly recent developments in qPCR markers allow researchers to explore the relative abundance of AM fungal species within roots in greenhouse experiments as well as in the field to identify local AM fungal communities.
qPCR markers for arbuscular mycorrhizal fungi will consist of AM specific primers and fluorescently labeled hydrolysis probes. These AM specific primers (discussed above) can be chosen by the researcher and this decision is typically guided by the question at hand, resources available, and willingness to troubleshoot in the lab.
Microarray
editDNA microarray analysis is currently being used in AM fungal research to simultaneously measure the expression of many genes from target species or experimental samples. The most common tool or method is to use functional gene array (FGA) technology, a specialized microarray that contains probes for genes that are functionally important in microbial processes such as carbon, nitrogen or phosphorus cycling. FGAs have the ability to simultaneously examine many functional genes.[96] This technique is typically used for general analysis of functional microbial genes, but when complemented with genetic sequencing, inferences can be made about the connection between fungal community composition and microbial functionality.
Specific organismal chemical signatures can be used to detect biomass of more cryptic organisms, such as AM fungi or soil bacteria. Lipids, more specifically phospholipids and neutral lipids, contain fatty acids connected to a glycerol backbone. The fatty acid composition of organisms varies, and the proportions of specific fatty acids can be organism specific. For example, in AM fungi the proportion of the fatty acids, 16:1ω5 and 18:1ω7, in the phospholipid portion account for approximately 58% of total fatty acid composition.[97] The fatty acid, 16:1ω5 is the most commonly used acid to characterize AM fungi in soils and can be used as a strong indicator of mycelial biomass in soil sample.[97]
Neutral lipid fatty acid analysis of AM fungi is typically looked upon as a method to indicate energy storage, but most importantly, the ratio of NLFA (16:1ω5) to PLFA (16:1ω5) can potentially be used to indicate nutritional status of AM fungi in soils. Energy is mainly stored in AM fungi as neutral lipids in storage structures like spores and vesicles. Because of this NLFA correlates quite well with the number of spores in a given volume of soil.[97] The ratio of NLFA concentration to PLFA concentration (active mycelia) can then give the proportion of carbon allocated to storage structures (spores, measured as NLFA).
Problems with lipid fatty acid analyses include the incomplete specificity of fatty acids to AM fungi, the species- or genera-specific variation in fatty acid composition can complicate analysis in systems with multiple AM fungal species (e.g. field soil), the high background levels of certain fatty acid concentration in soils, and that phospholipids are correlated to an organism's membrane area, and the surface to volume ratio can vary widely between organisms such as bacteria and fungi.[98] More work must be done to identify the efficacy of this method in field soils with many genera and species of AM fungi to discern the methods ability to discriminate between many varying fatty acid compositions.
Future research directions with AM fungi
editOne prospect for future analysis of AM fungi is the use of stable isotope probes. Stable isotope probing (SIP) is a technique that can be used to determine the active metabolic function of individual taxa within a complex system of microbes. This level of specificity, linking microbial function and phylogenetics, has not been achieved previously in microbial ecology. This method can also be used independently of classical culture methods in microbial ecology, allowing for in situ analysis of functional microbes. Application of sequencing of single nucleus from spores of AM fungi has also been developed recently and also circumvents the need of culture methods.[99]
Stable Isotope Probing (SIP)
editSIP, more explicitly DNA/RNA-based SIP, uses stable-isotope enriched substrates, such as 13C, 15N, or H218O, and then analyzes the 'labeled' markers using species specific DNA or RNA markers.[100] The analysis of labeled DNA is performed by separating unlabeled and labeled DNA on a cesium chloride gradient formed in an ultra centrifuge.[101] Because all microbial organisms are capable of importing water into their cells, the use of H218O stable isotope probing is a very exciting new method that can shed light on questions microbial ecologists and biologists have struggled with answering for years, in particular, what are the active microbial organisms in my system? The H218O, or heavy water method will target all organisms that are actively growing, and induce little influence on growth itself. This would be especially true with most greenhouse experiments with arbuscular mycorrhizas because plants must be watered anyway, and water does not directly select for organisms with specific metabolic pathways,[101] as would happen when using 13C and15N.
Little has been done with this method in arbuscular mycorrhizal experiments, but if proven to work in a controlled experiment, and with further refinement of DNA/RNA fungal community analyses techniques, this may be a viable option to very specifically determine the actively growing portion of AM fungal species across growing seasons, with different plant hosts or treatments, and in the face of climate change.
sRNA and sRNA processing mechanism to understand AM symbiosis
editsRNAs have been reported to take crucial role in the crosstalk between host and symbiont.[102] sRNAs processing mechanism is thus, important for understanding AM symbiosis. It seems that AM fungi have their unique features to have bacterial type core enzyme as well as the large number of Argonaute proteins in their sRNA processing system (or RNAi system).[17] sRNA and sRNA processing mechanism research is also exciting topic to understand AM fungi symbiosis.
Phytoremediation
editDisturbance of native plant communities in desertification-threatened areas is often followed by degradation of physical and biological soil properties, soil structure, nutrient availability, and organic matter. When restoring disturbed land, it is essential to replace not only the above ground vegetation but also biological and physical soil properties.[103]
A relatively new approach to restoring land is to inoculate soil with AM fungi when reintroducing vegetation in ecological restoration projects (phytoremediation). It has enabled host plants to establish themselves on degraded soil and improve soil quality and health.[104] Soils' quality parameters were significantly improved long-term when a mixture of indigenous arbuscular mycorrhizal fungi species was introduced compared to noninoculated soil and soil inoculated with a single exotic species of AM fungi.[103] The benefits were increased plant growth, increased phosphorus uptake[105] and soil nitrogen content, higher soil organic matter content, and soil aggregation, attributed to higher legume nodulation in the presence of AM fungi, better water infiltration, and soil aeration due to soil aggregation.[103] Native strains of AM fungi enhance the extraction of heavy metal(s) from the polluted soils and make the soil healthy and suitable for crop production.[106]
Agriculture
editon plant performance and soil fertility
On the right: mycorrhizal helper (MH) and plant growth promoting (PGP) bacteria synergistically interacting with AMF.[107]
Many modern agronomic practices are disruptive to mycorrhizal symbiosis. There is great potential for low-input agriculture to manage the system in a way that promotes mycorrhizal symbiosis.
Conventional agriculture practices, such as tillage, heavy fertilizers and fungicides, poor crop rotations, and selection for plants that survive these conditions, hinder the ability of plants to form symbiosis with arbuscular mycorrhizal fungi.
Most agricultural crops can perform better and are more productive when well-colonized by AM fungi. AM symbiosis increases the phosphorus and micronutrient uptake and growth of their plant host (George et al. 1992).
Management of AM fungi is especially important for organic and low-input agriculture systems where soil phosphorus is, in general, low, although all agroecosystems can benefit by promoting arbuscular mycorrhizae establishment.
Some crops that are poor at seeking out nutrients in the soil are very dependent on AM fungi for phosphorus uptake. For example, flax, which has poor chemotaxic ability, is highly dependent on AM-mediated phosphorus uptake at low and intermediate soil phosphorus concentrations (Thingstrup et al. 1998).
Proper management of AMF in the agroecosystems can improve the quality of the soil and the productivity of the land. Agricultural practices such as reduced tillage, low phosphorus fertilizer usage, and perennialized cropping systems promote functional mycorrhizal symbiosis.
Tillage
editTillage reduces the inoculation potential of the soil and the efficacy of mycorrhizaes by disrupting the extraradical hyphal network (Miller et al. 1995, McGonigle & Miller 1999, Mozafar et al. 2000).
By breaking apart the soil macro structure, the hyphal network is rendered non-infective (Miller et al. 1995, McGonigle & Miller 1999). The disruption of the hyphal network decreases the absorptive abilities of the mycorrhizae because the surface area spanned by the hyphae is greatly reduced. This, in turn, lowers the phosphorus input to the plants that are connected to the hyphal network (Figure 3, McGonigle & Miller 1999).
In reduced-tillage system, heavy phosphorus fertilizer input may not be required as compared to heavy-tillage systems. This is due to the increase in mycorrhizal network, which allows mycorrhizae to provide the plant with sufficient phosphorus (Miller et al. 1995).
Phosphorus fertilizer
editThe benefits of AMF are greatest in systems where inputs are low. Heavy usage of phosphorus fertilizer can inhibit mycorrhizal colonization and growth.
As the soil's phosphorus levels available to the plants increases, the amount of phosphorus also increases in the plant's tissues, and carbon drain on the plant by the AM fungi symbiosis become non-beneficial to the plant (Grant 2005).
A decrease in mycorrhizal colonization due to high soil-phosphorus levels can lead to plant deficiencies in other micronutrients that have mycorrhizal-mediated uptake such as copper (Timmer & Leyden 1980).
Perennialized cropping systems
editCover crops are grown in the fall, winter, and spring, covering the soil during periods when it would commonly be left without a cover of growing plants.
Mycorrhizal cover crops can be used to improve the mycorrhizal inoculum potential and hyphal network (Kabir and Koide 2000, Boswell et al.1998, Sorensen et al. 2005).
Since AM fungi are biotrophic, they are dependent on plants for the growth of their hyphal networks. Growing a cover crop extends the time for AM growth into the autumn, winter, and spring. Promotion of hyphal growth creates a more extensive hyphal network. The mycorrhizal colonization increase found in cover crops systems may be largely attributed to an increase in the extraradical hyphal network that can colonize the roots of the new crop (Boswell et al. 1998). The extraradical mycelia are able to survive the winter, providing rapid spring colonization and early season symbiosis (McGonigle and Miller 1999). This early symbiosis allows plants to tap into the well-established hyphal network and be supplied with adequate phosphorus nutrition during early growth, which greatly improves the crop yield.
Soil quality
editRestoration of native AM fungi increases the success of ecological restoration projects and the rapidity of soil recovery.[103] AM fungi enhance soil aggregate stability by the production of extraradical hyphae and a soil protein known as glomalin.
Glomalin-related soil proteins (GRSP) have been identified using a monoclonal antibody (Mab32B11) raised against crushed AMF spores. It is defined by its extraction conditions and reaction with the antibody Mab32B11.
There is other circumstantial evidence to show that glomalin is of AM fungal origin. When AM fungi are eliminated from soil through incubation of soil without host plants, the concentration of GRSP declines. A similar decline in GRSP has also been observed in incubated soils from forested, afforested, and agricultural land[108] and grasslands treated with fungicide.[109]
Glomalin is hypothesized to improve soil aggregate water stability and decrease soil erosion. A strong correlation has been found between GRSP and soil aggregate water stability in a wide variety of soils where organic material is the main binding agent, although the mechanism is not known.[109] The protein glomalin has not yet been isolated and described, and the link between glomalin, GRSP, and arbuscular mycorrhizal fungi is not yet clear.[109]
See also
editReferences
edit- ^ Mycorrhizal Symbiosis. 2008. doi:10.1016/b978-0-12-370526-6.x5001-6. ISBN 978-0-12-370526-6.
- ^ Spatafora, Joseph W.; Chang, Ying; Benny, Gerald L.; Lazarus, Katy; Smith, Matthew E.; Berbee, Mary L.; Bonito, Gregory; Corradi, Nicolas; Grigoriev, Igor; Gryganskyi, Andrii; James, Timothy Y.; O'Donnell, Kerry; Roberson, Robert W.; Taylor, Thomas N.; Uehling, Jessie (2016). "A phylum-level phylogenetic classification of zygomycete fungi based on genome-scale data". Mycologia. 108 (5): 1028–1046. doi:10.3852/16-042. ISSN 0027-5514. PMC 6078412. PMID 27738200.
- ^ Brundrett, M.C. (2002). "Coevolution of roots and mycorrhizas of land plants". New Phytologist. 154 (2): 275–304. doi:10.1046/j.1469-8137.2002.00397.x. PMID 33873429.
- ^ Barbour, M.G.; Burk, J.H.; Pitts, W.D. (1980). Terrestrial plant ecology. Frontiers in Physics. Benjamin/Cummings Publishing Company. p. 118. ISBN 978-0-8053-0540-1.
- ^ a b Simon, L.; Bousquet, J.; Levesque, C.; Lalonde, M. (1993). "Origin and diversification of endomycorrhizal fungi and coincidence with vascular land plants". Nature. 363 (6424): 67–69. Bibcode:1993Natur.363...67S. doi:10.1038/363067a0. S2CID 4319766.
- ^ Schüßler, A.; et al. (2001). "A new fungal phylum, the Glomeromycota: phylogeny and evolution". Mycol. Res. 105 (12): 1413–1421. doi:10.1017/S0953756201005196.
- ^ Moore, David; Robson, Geoffrey D.; Trinci, Anthony P. J. (2020), 21st Century Guidebook to Fungi, doi:10.1017/9781108776387, ISBN 9781108776387
- ^ Smith, Sally A; Read, David (2008), "2 - Colonization of roots and anatomy of arbuscular mycorrhizas", Mycorrhizal Symbiosis (Third ed.): 42–90, doi:10.1016/B978-012370526-6.50004-0, ISBN 9780123705266
- ^ Jacott, Catherine; Murray, Jeremy; Ridout, Christopher (2017). "Trade-Offs in Arbuscular Mycorrhizal Symbiosis: Disease Resistance, Growth Responses and Perspectives for Crop Breeding". Agronomy. 7 (4): 75. doi:10.3390/agronomy7040075. This article incorporates text from this source, which is available under the CC BY 4.0 license.
- ^ a b c d e Remy, W.; Taylor, T.; Hass, H.; Kerp, H. (1994). "Four hundred-million-year-old vesicular arbuscular mycorrhizae". Proceedings of the National Academy of Sciences of the United States of America. 91 (25): 11841–11843. Bibcode:1994PNAS...9111841R. doi:10.1073/pnas.91.25.11841. PMC 45331. PMID 11607500.
- ^ Kar, R.K.; Mandaokar, B.D.; Kar, R. (2005). "Mycorrhizal fossil fungi from the Miocene sediments of Mirozam, Northeast India". Current Science. 89: 257–259.
- ^ Strullu-Derrien, Christine; Strullu, Désiré-Georges (November 2007), "Mycorrhization of fossil and living plantsLa mycorrhization des plantes fossiles et actuelles", Comptes Rendus Palevol, La paléobotanique et l'évolution du monde végétal : Quelques problèmes d'actualité, 6 (6–7): 483–494, doi:10.1016/j.crpv.2007.09.006
- ^ a b Krings, Michael; Harper, Carla J; Taylor, Edith L. (2018), "Fungi and fungal interactions in the Rhynie chert: a review of the evidence, with the description of Perexiflasca tayloriana gen. et sp. nov", Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences, 373 (1739), doi:10.1098/rstb.2016.0500, PMC 5745336, PMID 29254965
- ^ Strullu-Derrien, Christine; Selosse, Marc-André; Kenrik, Paul; Martin, Francis M. (24 March 2018), "The origin and evolution of mycorrhizal symbioses: from palaeomycology to phylogenomics", New Phytologist, 220 (4): 1012–1030, doi:10.1111/nph.15076, ISSN 1469-8137, PMID 29573278
- ^ a b Wang, B.; Yeun, L.H.; Xue, Y.; Liu, Y.; Ane, J.M.; Qiu, Y.L. (2010). "Presence of three mycorrhizal genes in the common ancestor of land plants suggests a key role of mycorrhizas in the colonization of land by plants". New Phytologist. 186 (2): 514–525. doi:10.1111/j.1469-8137.2009.03137.x. hdl:2027.42/78704. PMID 20059702.
- ^ Ané, JM; et al. (November 2002). "Genetic and cytogenetic mapping of DMI1, DMI2, and DMI3 genes of Medicago truncatula involved in Nod factor transduction, nodulation, and mycorrhization". Molecular Plant-Microbe Interactions. 15 (11): 1108–18. doi:10.1094/MPMI.2002.15.11.1108. PMID 12423016.
- ^ a b Lee, Soon-Jae; Kong, Mengxuan; Harrison, Paul; Hijri, Mohamed (2018), "Conserved Proteins of the RNA Interference System in the Arbuscular Mycorrhizal Fungus Rhizoglomus irregulare Provide New Insight into the Evolutionary History of Glomeromycota", Genome Biology and Evolution, 10 (1): 328–343, doi:10.1093/gbe/evy002, PMC 5786227, PMID 29329439
- ^ Alexopolous C, Mims C, Blackwell M. 2004. Introductory mycology, 4th ed . Hoboken (NJ): John Wiley and Sons.
- ^ a b Lee, SJ., Kong, M., Morse, D. et al. Expression of putative circadian clock components in the arbuscular mycorrhizal fungus Rhizoglomus irregulare. Mycorrhiza (2018) https://doi.org/10.1007/s00572-018-0843-y
- ^ Marleau J; Dalpé Y; St-Arnaud M; Hijri M (2011). "Spore development and nuclear inheritance in arbuscular mycorrhizal fungi". BMC Evolutionary Biology. 11 (1): 51. Bibcode:2011BMCEE..11...51M. doi:10.1186/1471-2148-11-51. PMC 3060866. PMID 21349193.
- ^ a b c d Wright S.F. (2005). "Management of Arbuscular Mycorrhizal Fungi". In R.W. Zobel; S.F. Wright (eds.). Roots and Soil Management: Interactions between roots and the soil. US: American Society of Agronomy. pp. 183–197.
- ^ a b c d Douds, D.D. and Nagahashi, G. 2000. Signalling and Recognition Events Prior to Colonisation of Roots by Arbuscular Mycorrhizal Fungi. In Current Advances in Mycorrhizae Research. Ed. Podila, G.K., Douds, D.D. Minnesota: APS Press. Pp 11–18.
- ^ Akiyama K; Matsuzaki K; Hayashi H (2005). "Plant sesquiterpenes induce hyphal branching in arbuscular mycorrhizal fungi". Nature. 435 (7043): 824–827. Bibcode:2005Natur.435..824A. doi:10.1038/nature03608. PMID 15944706. S2CID 4343708.
- ^ a b c d Nagahashi, G; Douds, D. D.; Abney, G.D. (1996). "Phosphorus amendment inhibits hyphal branching of VAM fungus Gigaspora margarita directly and indirectly through its effect on root exudation". Mycorrhiza. 6 (5): 403–408. Bibcode:1996Mycor...6..403N. doi:10.1007/s005720050139. S2CID 36014515.
- ^ Sbrana, C.; Giovannetti, M. (2005). "Chemotropism in the arbuscular mycorrhizal fungus Glomus mosseae". Mycorrhiza. 15 (7): 539–545. Bibcode:2005Mycor..15..539S. doi:10.1007/s00572-005-0362-5. PMID 16133246. S2CID 23648484.
- ^ Tamasloukht, M.; Sejalon-Delmas, N.; Kluever, A.; Jauneau, A.; Roux., C.; Becard, G.; Franken, P. (2003). "Root Factors Induce Mitochondrial-Related Gene Expression and Fungal Respiration during the Developmental Switch from Asymbiosis to Presymbiosis in the Arbuscular Mycorrhizal Fungus Gigaspora rosea". Plant Physiology. 131 (3): 1468–1478. doi:10.1104/pp.012898. PMC 166906. PMID 12644696.
- ^ a b c Gianinazzi-Pearson, V. (1996). "Plant Cell Responses to Arbuscular Mycorrhizal Fungi: Getting to the Roots of the Symbiosis". The Plant Cell. 8 (10): 1871–1883. doi:10.1105/tpc.8.10.1871. JSTOR 3870236. PMC 161321. PMID 12239368.
- ^ Lara Armstrong; R. Larry Peterson; Lara Armstrong; R. Larry Peterson (2002). "The Interface between the Arbuscular Mycorrhizal Fungus Glomus intraradices and Root Cells of Panax quinquefolius: A Paris-Type Mycorrhizal Association". Mycologia. 94 (4): 587–595. doi:10.2307/3761710. JSTOR 3761710. PMID 21156532.
- ^ Yamato, Masahide (2005). "Morphological types of arbuscular mycorrhizas in pioneer woody plants growing in an oil palm farm in Sumatra, Indonesia". Mycoscience. 46: 66–68. doi:10.1007/s10267-004-0212-x. S2CID 83767351.
- ^ Matekwor, Ahulu E; Nakata, M; Nonaka, M (Mar 2005). "Arum- and Paris-type arbuscular mycorrhizas in a mixed pine forest on sand dune soil in Niigata Prefecture, central Honshu, Japan". Mycorrhiza. 15 (2): 129–36. Bibcode:2005Mycor..15..129M. doi:10.1007/s00572-004-0310-9. ISSN 0940-6360. PMID 15290409. S2CID 25476630.
- ^ Tuomi, J.; Kytoviita, M.; Hardling, R. (2001). "Cost efficiency of nutrient acquisition of mycorrhizal symbiosis for the host plant". Oikos. 92: 62–70. doi:10.1034/j.1600-0706.2001.920108.x.
- ^ a b c Bolan, N.S. (1991). "A critical review of the role of mycorrhizal fungi in the uptake of phosphorus by plants". Plant and Soil. 134 (2): 189–207. Bibcode:1991PlSoi.134..189B. doi:10.1007/BF00012037. S2CID 44215263.
- ^ Harley, J.L., Smith, S.E., 1983. Mycorrhizal Symbiosis. Academic Press: London.
- ^ a b c d Pfeffer, P.; Douds D.; Becard, G.; Shachar-Hill, Y. (1999). "Carbon Uptake and the Metabolism and Transport of Lipids in an Arbuscular Mycorrhiza". Plant Physiology. 120 (2): 587–598. doi:10.1104/pp.120.2.587. PMC 59298. PMID 10364411.
- ^ a b c Hamel, C. (2004). "Impact of arbuscular mycorrhiza fungi on N and P cycling in the root zone". Canadian Journal of Soil Science. 84 (4): 383–395. doi:10.4141/S04-004.
- ^ H. Bücking and Y.Shachar-Hill (2005). Phosphate uptake, transport and transfer by the arbuscular mycorrhizal fungus Glomus intraradices is stimulated by increased carbohydrate availability New Phytologist 165:899–912
- ^ a b c Smith, S.; Smith, A.; Jakobsen, I. (2003). "Mycorrhizal Fungi Can Dominate Phosphate Supply to Plants Irrespective of Growth Responses". Plant Physiology. 133 (1): 16–20. doi:10.1104/pp.103.024380. PMC 1540331. PMID 12970469.
- ^ a b Mateus, I. D.; Auxier, B.; Ndiaye MMS; Cruz, J.; Lee, S. J.; Sanders, I. R. (2022). "Reciprocal recombination genomic signatures in the symbiotic arbuscular mycorrhizal fungi Rhizophagus irregularis". PLOS ONE. 17 (7): e0270481. Bibcode:2022PLoSO..1770481M. doi:10.1371/journal.pone.0270481. PMC 9249182. PMID 35776745.
- ^ Halary, Sébastien; Malik, Shehre-Banoo; Lildhar, Levannia; Slamovits, Claudio H.; Hijri, Mohamed; Corradi, Nicolas (2011). "Conserved Meiotic Machinery in Glomus SPP., a Putatively Ancient Asexual Fungal Lineage". Genome Biology and Evolution. 3: 950–958. doi:10.1093/gbe/evr089. PMC 3184777. PMID 21876220.
- ^ Maillet, Fabienne; Poinsot, Véréna; André, Olivier; Puech-Pagès, Virginie; Haouy, Alexandra; Gueunier, Monique; Cromer, Laurence; Giraudet, Delphine; Formey, Damien; Niebel, Andreas; Martinez, Eduardo Andres; Driguez, Hugues; Bécard, Guillaume; Dénarié, Jean (January 2011). "Fungal lipochitooligosaccharide symbiotic signals in arbuscular mycorrhiza". Nature. 469 (7328): 58–63. Bibcode:2011Natur.469...58M. doi:10.1038/nature09622. PMID 21209659. S2CID 4373531.
- ^ Cope, Kevin R.; Bascaules, Adeline; Irving, Thomas B.; Venkateshwaran, Muthusubramanian; Maeda, Junko; Garcia, Kevin; Rush, Tomás A.; Ma, Cathleen; Labbé, Jessy; Jawdy, Sara; Steigerwald, Edward; Setzke, Jonathan; Fung, Emmeline; Schnell, Kimberly G.; Wang, Yunqian; Schleif, Nathaniel; Bücking, Heike; Strauss, Steven H.; Maillet, Fabienne; Jargeat, Patricia; Bécard, Guillaume; Puech-Pagès, Virginie; Ané, Jean-Michel (October 2019). "The Ectomycorrhizal Fungus Laccaria bicolor Produces Lipochitooligosaccharides and Uses the Common Symbiosis Pathway to Colonize Populus Roots". The Plant Cell. 31 (10): 2386–2410. doi:10.1105/tpc.18.00676. PMC 6790088. PMID 31416823.
- ^ MacLean, Allyson M.; Bravo, Armando; Harrison, Maria J. (October 2017). "Plant Signaling and Metabolic Pathways Enabling Arbuscular Mycorrhizal Symbiosis". The Plant Cell. 29 (10): 2319–2335. doi:10.1105/tpc.17.00555. PMC 5940448. PMID 28855333.
- ^ a b c d e f g Smith, S.E., Read D.J. Mycorrhizal Symbiosis. 2002. Academic Press: London.
- ^ a b Smith, Read, Sally, DJ (2008). Mycorrhizal symbiosis. New York: Academic Press.
{{cite book}}: CS1 maint: multiple names: authors list (link) - ^ a b c Opik, M; Vanatoa A; Vanatoa E; Moora M; Davidson J; Kalwij JM; Reier U; Zobel M (2010). "The online database MaarjAM reveals global and ecosystemic distribution patterns in arbuscular mycorrhizal fungi (Glomeromycota)". New Phytologist. 188 (1): 233–241. doi:10.1111/j.1469-8137.2010.03334.x. PMID 20561207.
- ^ a b Kivlin, Stephanie; Christine V. Hawkes; Kathleen K. Treseder (2011). "Global diversity and distribution of arbuscular mycorrhizal fungi". Soil Biology and Biochemistry. 43 (11): 2294–2303. Bibcode:2011SBiBi..43.2294K. doi:10.1016/j.soilbio.2011.07.012. S2CID 85135958.
- ^ Lekberg, Y; Koide RT; Rohr JR; Aldirch-Wolfe L; Morton JB (2007). "Role of niche restrictions and dispersal in the composition of arbuscular mycorrhizal fungal communities". Journal of Ecology. 95 (1): 95–100. Bibcode:2007JEcol..95...95L. doi:10.1111/j.1365-2745.2006.01193.x.
- ^ Leff, Jonathan W.; Jones, Stuart E.; Prober, Suzanne M.; Barberán, Albert; Borer, Elizabeth T.; Firn, Jennifer L.; Harpole, W. Stanley; Hobbie, Sarah E.; Hofmockel, Kirsten S. (2015-09-01). "Consistent responses of soil microbial communities to elevated nutrient inputs in grasslands across the globe". Proceedings of the National Academy of Sciences. 112 (35): 10967–10972. Bibcode:2015PNAS..11210967L. doi:10.1073/pnas.1508382112. ISSN 0027-8424. PMC 4568213. PMID 26283343.
- ^ Allen, EB; Allen MF; Helm DJ; Trappe JM; Molina R; Rincon E (1995). "Patterns and regulation of mycorrhizal plant and fungal diversity". Plant and Soil. 170 (1): 47–62. Bibcode:1995PlSoi.170...47A. doi:10.1007/bf02183054. S2CID 25097125.
- ^ Klironomos, John (2000). Host-specificity and functional diversity among arbuscular mycorrhizal fungi. Halifax, Canada: Microbial Biosystems: New Frontiers. Proceedings of the 8th International Symposium on Microbial Ecology. Atlantic Canada Society for Microbial Ecology. pp. 845–851.
- ^ Husband, R; Herre EA; Turner SL; Gallery R; Young JPW (2002). "Molecular diversity of arbuscular mycorrhizal fungi and patterns of associations over time and space in a tropical forest" (PDF). Molecular Ecology. 11 (12): 2669–2678. doi:10.1046/j.1365-294x.2002.01647.x. PMID 12453249. S2CID 10279037.
- ^ a b Hempel, Stefan; Gotzenberger L; Kuhn I; Michalski SG; Rillig M; Zobel M; Moora M (2013). "Mycorrhizas in the Central European flora – relationships with plant life history traits and ecology". Ecology. 94 (6): 1389–1399. Bibcode:2013Ecol...94.1389H. doi:10.1890/12-1700.1. PMID 23923502.
- ^ Drigo, B; Pijl, AS; Duyts, H; Kielak, AM; Gamper, HA; Houtekamer, MJ; Boschker, HTS; Bodelier, PLE; Whiteley, AS; Veen, JAV; Kowalchuk, GA (2010). "Shifting carbon flow from roots into associated microbial communities in response to elevated atmospheric CO2". Proceedings of the National Academy of Sciences of the United States of America. 107 (24): 10938–10942. Bibcode:2010PNAS..10710938D. doi:10.1073/pnas.0912421107. PMC 2890735. PMID 20534474.
- ^ van der Heijden, MG; Boller AT; Wiemken A; Sanders IR (1998). "Different arbuscular mycorrhizal fungi species are potential determinants of plant community structure". Ecology. 79 (6): 2082–2091. doi:10.1890/0012-9658(1998)079[2082:damfsa]2.0.co;2.
- ^ van der Heijden, MGA; Bardgett RD; Van Straalen NM (2008). "The unseen majority: soil microbes as drivers of plant diversity and productivity in terrestrial ecosystems". Ecology Letters. 11 (3): 296–310. Bibcode:2008EcolL..11..296V. doi:10.1111/j.1461-0248.2007.01139.x. PMID 18047587.
- ^ Vannette, RL; Rasmann S (2012). "Arbuscular mycorrhizal fungi mediate below-ground plant–herbivore interactions: a phylogenetic study". Functional Ecology. 26 (5): 1033–1042. Bibcode:2012FuEco..26.1033V. doi:10.1111/j.1365-2435.2012.02046.x.
- ^ Haumann, N; Hawkes CV (2009). "Plant neighborhood control of arbuscular mycorrhizal community composition". New Phytologist. 183 (4): 1188–1200. doi:10.1111/j.1469-8137.2009.02882.x. PMID 19496954.
- ^ Hausmann, N; Hawkes CV (2010). "Order of plant host establishment alters the composition of arbuscular mycorrhizal communities". Ecology. 91 (8): 2333–23343. Bibcode:2010Ecol...91.2333H. doi:10.1890/09-0924.1. PMID 20836455.
- ^ Batten, KM; Skow KM; Davies KF; Harrison SP (2006). "Two invasive plants alter soil microbial community composition in serpentine grasslands". Biological Invasions. 8 (2): 217–230. Bibcode:2006BiInv...8..217B. doi:10.1007/s10530-004-3856-8. S2CID 24969103.
- ^ Hawkes, CV; Belnap J; D'Antonio C; Firestone M (2006). "Arbuscular mycorrhizal assemblages in native plant roots change in the presence of invasive exotic grasses". Plant and Soil. 281 (1–2): 369–380. Bibcode:2006PlSoi.281..369H. doi:10.1007/s11104-005-4826-3. S2CID 16801874.
- ^ Kivlin, Stephanie; Christine V. Hawkes (2011). "Differentiating between effects of invasion and diversity: impacts of aboveground plant communities on belowground fungal communities". New Phytologist. 189 (2): 526–535. doi:10.1111/j.1469-8137.2010.03494.x. PMID 20958304.
- ^ Lekberg, Y; Gibbons SM; Rosendahl S; Ramsey PW (2013). "Severe plant invasions can increase mycorrhizal fungal abundance and diversity". ISME Journal. 7 (7): 1424–1433. Bibcode:2013ISMEJ...7.1424L. doi:10.1038/ismej.2013.41. PMC 3695300. PMID 23486251.
- ^ Fitter, AH (2005). "Darkness visible: reflections on underground ecology" (PDF). Journal of Ecology. 93 (2): 231–243. Bibcode:2005JEcol..93..231F. doi:10.1111/j.0022-0477.2005.00990.x.
- ^ Pringle, A; Bever, JD; Gardes, M; Parrent, JL; Rillig, MC; Klironomos, JN (2009). "Mycorrhizal symbioses and plant invasions". Annual Review of Ecology, Evolution, and Systematics. 40: 699–715. doi:10.1146/annurev.ecolsys.39.110707.173454.
- ^ Larimer, AL; Bever JD; Clay K (2010). "The interactive effects of plant microbial symbionts: a review and meta-analysis". Symbiosis. 51 (2): 139–148. Bibcode:2010Symbi..51..139L. doi:10.1007/s13199-010-0083-1. S2CID 11569239.
- ^ Novas, MV; Iannone LJ; Godeas AM; Cabral D (2009). "Positive association between mycorrhiza and foliar endophytes in a Poa bonariensis". Mycological Progress. 8: 75–81. doi:10.1007/s11557-008-0579-8. hdl:11336/27622. S2CID 2206235.
- ^ Larimer, AL; Bever JD; Clay K (2012). "Consequences of simultaneous interactions of fungal endophytes and arbuscular mycorrhizal fungi with a shared host grass". Oikos. 121 (12): 2090–2096. Bibcode:2012Oikos.121.2090L. doi:10.1111/j.1600-0706.2012.20153.x.
- ^ Omacini, M; Eggers T; Bonkowski M; Gange AC; Jones TH (2006). "Leaf endophytes affect mycorrhizal status and growth of co-infected and neighboring plants". Functional Ecology. 20 (2): 226–232. Bibcode:2006FuEco..20..226O. doi:10.1111/j.1365-2435.2006.01099.x.
- ^ Mack, KML; Rudgers JA (2008). "Balancing multiple mutualists: asymmetric interactions among plants, arbuscular mycorrhizal fungi, and fungal endophytes". Oikos. 117 (2): 310–320. Bibcode:2008Oikos.117..310M. CiteSeerX 10.1.1.722.4169. doi:10.1111/j.2007.0030-1299.15973.x.
- ^ Liu, QH; Parsons AJ; Xue H; Fraser K; Ryan GD; Newman JA; Rasmussen S (2011). "Competition between foliar Neotyphodium lolii endophytes and mycorrhizal Glomus spp. fungi in Lolium perenne depends on resource supply and host carbohydrate content". Functional Ecology. 25 (4): 910–920. Bibcode:2011FuEco..25..910L. doi:10.1111/j.1365-2435.2011.01853.x. hdl:10214/3316.
- ^ Reininger, V; Sieber TN (2012). "Mycorrhiza reduces adverse effects of dark septate endophytes (DSE) on growth of conifers". PLOS ONE. 7 (8): 1–10. Bibcode:2012PLoSO...742865R. doi:10.1371/journal.pone.0042865. PMC 3416760. PMID 22900058.
- ^ Klironomos, JN; Hart MM; Gurney JE; Moutoglis P (2001). "Interspecific differences in the tolerance of arbuscular mycorrhizal fungi to freezing and drying". Canadian Journal of Botany. 79 (10): 1161–1166. doi:10.1139/cjb-79-10-1161.
- ^ Auge, RM (2001). "Water relations, drought and vesicular-arbuscular mycorrhizal symbiosis". Mycorrhiza. 11 (1): 3–42. Bibcode:2001Mycor..11....3A. doi:10.1007/s005720100097. S2CID 29379395.
- ^ Johnson, NC; Rowland DL; Corkidi L; Egerton-Warburton LM; Allen EB (2003). "Nitrogen enrichment alters mycorrhizal allocation at five mesic to semiarid grasslands". Ecology. 84 (7): 1895–1908. doi:10.1890/0012-9658(2003)084[1895:neamaa]2.0.co;2.
- ^ Johnson, NC; Wilson GWT; Bowker MA; Wilson JA; Miller RM (2010). "Resource limitation is a driver of local adaptation in mycorrhizal symbioses". Proceedings of the National Academy of Sciences of the United States of America. 107 (5): 2093–2098. Bibcode:2010PNAS..107.2093J. doi:10.1073/pnas.0906710107. PMC 2836645. PMID 20133855.
- ^ Gai, JP; Tian H; Yang FY; Christie P; Li XL; Klironomos JN (2012). "Arbuscular mycorrhizal fungal diversity along a Tibetan elevation gradient". Pedobiologia. 55 (3): 145–151. Bibcode:2012Pedob..55..145G. doi:10.1016/j.pedobi.2011.12.004.
- ^ Porcel, R; Aroca, R; Ruiz-Lozano, JM (2012). "stress alleviation using arbuscular mycorrhizal fungi. A review" (PDF). Agronomy for Sustainable Development. 32: 181–200. doi:10.1007/s13593-011-0029-x. S2CID 8572482.
- ^ a b c Marschner, P.; Timonen, S. (2004). "Interactions between plant species and mycorrhizal colonization on the bacterial community composition in the rhizosphere". Applied Soil Ecology. 28: 23–36. doi:10.1016/j.apsoil.2004.06.007.
- ^ Eriksson, A. (2001). "Arbuscular mycorrhizae in relation to management history, soil nutrients and plant diversity". Plant Ecology. 155 (2): 129–137. doi:10.1023/A:1013204803560. S2CID 42097761.
- ^ "Lipochitooligosaccharides Stimulating Arbuscular Mycorrhizal Symbiosis". Archived from the original on 2012-08-05.
- ^ Schrey, Silvia D.; et al. (2012). "Production of fungal and bacterial growth modulating secondary metabolites is widespread among mycorrhiza-associated streptomycetes". BMC Microbiology. 12 (1): 164. doi:10.1186/1471-2180-12-164. PMC 3487804. PMID 22852578.
- ^ Van der Putten, WH (2012). "Climate change, Aboveground-belowground interactions and species' range shifts". Annual Review of Ecology, Evolution, and Systematics. 43: 365–383. doi:10.1146/annurev-ecolsys-110411-160423. hdl:20.500.11755/0f7d61b9-e435-4da3-b3d6-3ca115bdbdb5. S2CID 85941864.
- ^ Worchel, Elise; Giauque, Hannah E.; Kivlin, Stephanie N. (2013). "Fungal symbionts alter plant drought response". Microbial Ecology. 65 (3): 671–678. Bibcode:2013MicEc..65..671W. doi:10.1007/s00248-012-0151-6. PMID 23250115. S2CID 122731.
- ^ Kivlin, SN; Emery SM; Rudgers JA (2013). "Fungal symbionts alter plant response to global change". American Journal of Botany. 100 (7): 1445–1457. doi:10.3732/ajb.1200558. PMID 23757444.
- ^ Treseder, KK (2004). "A meta-analysis of mycorrhizal responses to nitrogen, phosphorus, and atmospheric CO2 in field studies". New Phytologist. 164 (2): 347–355. doi:10.1111/j.1469-8137.2004.01159.x. PMID 33873547.
- ^ Glomalin. "Hiding Place for a Third of the World's Stored Soil Carbon". Agricultural Research Journal.
- ^ Stukenbrock, Eva; Rosendahl, Soren (2005). "Clonal diversity and population genetic structure of arbuscular mycorrhizal fungi ( Glomus spp.) studied by multilocus genotyping of single spores". Molecular Ecology. 14 (3): 743–752. Bibcode:2005MolEc..14..743S. doi:10.1111/j.1365-294x.2005.02453.x. PMID 15723666. S2CID 30799196.
- ^ Baoming, Ji; et al. (2013). "Patterns of diversity and adaptation in Glomeromycota from three prairie grasslands". Molecular Ecology. 22 (9): 2573–2587. Bibcode:2013MolEc..22.2573J. doi:10.1111/mec.12268. PMID 23458035. S2CID 6769986.
- ^ Opik, Maarja; et al. (2013). "Global sampling of plant roots expands the described molecular diversity of arbuscular mycorrhizal fungi". Mycorrhiza. 23 (5): 411–430. Bibcode:2013Mycor..23..411O. doi:10.1007/s00572-013-0482-2. PMID 23422950. S2CID 17162006.
- ^ Johnson, Nancy; Zak, D.R.; Tilman, D.; Pfleger, F.L. (1991). "Dynamics of vesicular arbuscular mycorrhizae during old field succession". Oecologia. 86 (3): 349–358. Bibcode:1991Oecol..86..349J. doi:10.1007/bf00317600. PMID 28312920. S2CID 31539360.
- ^ Marleau, Julie; Dalpé, Y.; St-Arnaud, M.; Hijri, M. (2011). "Spore development and nuclear inheritance in arbuscular mycorrhizal fungi". BMC Evolutionary Biology. 11 (1): 51. Bibcode:2011BMCEE..11...51M. doi:10.1186/1471-2148-11-51. PMC 3060866. PMID 21349193.
- ^ Ehinger, Martine O.; Daniel Croll; Alexander M. Koch; Ian R. Sanders (2012). "Significant genetic and phenotypic changes arising from clonal growth of a single spore of an arbuscular mycorrhizal fungus over multiple generations". New Phytologist. 196 (3): 853–861. doi:10.1111/j.1469-8137.2012.04278.x. PMID 22931497.
- ^ Simon, L; Lalonde, M.; Bruns, T.D (1992). "Specific amplification of 18S fungal ribosomal genes from vesicular-arbuscular endomycorrhizal fungi colonizing roots". Appl. Environ. Microbiol. 58 (1): 291–295. Bibcode:1992ApEnM..58..291S. doi:10.1128/AEM.58.1.291-295.1992. PMC 195206. PMID 1339260.
- ^ Helgason, Thorunn; Fitter, A.H.; Young, J.P.W (1999). "Molecular diversity of arbuscular mycorrhizal fungi colonising Hyacinthoides non-scripta (bluebell) in a seminatural woodland" (PDF). Molecular Ecology. 8 (4): 659–666. Bibcode:1999MolEc...8..659H. doi:10.1046/j.1365-294x.1999.00604.x. S2CID 85991904.
- ^ Redecker, Dirk; Arthur Schüßler; Herbert Stockinger; Sidney L. Stürmer; Joseph B. Morton; Christopher Walker (2013). "An evidence-based consensus for the classification of arbuscular mycorrhizal fungi (Glomeromycota)". Mycorrhiza. 23 (7): 515–531. Bibcode:2013Mycor..23..515R. doi:10.1007/s00572-013-0486-y. PMID 23558516. S2CID 16495856.
- ^ He, Zhili; Joy D. VAN NOSTRAND; Ye DENG; Jizhong ZHOU (2011). "Development and applications of functional gene microarrays in the analysis of the functional diversity, composition, and structure of microbial communities". Front. Environ. Sci. Eng. China. 5 (1): 1–20. doi:10.1007/s11783-011-0301-y.
- ^ a b c Olsson, Pal Axel (1999). "Signature fatty acids provide tools for determination of the distribution and interactions of mycorrhizal fungi in soil". FEMS Microbiology Ecology. 29 (4): 303–310. Bibcode:1999FEMME..29..303O. doi:10.1111/j.1574-6941.1999.tb00621.x.
- ^ J., Schnürer; Clarholm, M.; Rosswall, T (1985). "Microbial biomass and activity in an agricultural soil with different organic matter contents". Soil Biology and Biochemistry. 17 (5): 611–618. Bibcode:1985SBiBi..17..611S. doi:10.1016/0038-0717(85)90036-7.
- ^ Montoliu-Nerin, Merce; Sánchez-García, Marisol; Bergin, Claudia; Grabherr, Manfred; Ellis, Barbara; Kutschera, Verena Esther; Kierczak, Marcin; Johannesson, Hanna; Rosling, Anna (2020-01-28). "Building de novo reference genome assemblies of complex eukaryotic microorganisms from single nuclei". Scientific Reports. 10 (1): 1303. Bibcode:2020NatSR..10.1303M. doi:10.1038/s41598-020-58025-3. ISSN 2045-2322. PMC 6987183. PMID 31992756.
- ^ Dumont, Marc G.; Murrell, Colin J. (2005). "Stable isotope probing – linking microbial identity to function". Nature Reviews Microbiology. 3 (6): 499–504. doi:10.1038/nrmicro1162. PMID 15886694. S2CID 24051877.
- ^ a b Schwartz, Egbert (2007). "Characterization of growing microorganisms in soil by stable isotope probing with H218O". Applied and Environmental Microbiology. 73 (8): 2541–2546. Bibcode:2007ApEnM..73.2541S. doi:10.1128/AEM.02021-06. PMC 1855593. PMID 17322324.
- ^ Formey D, et al. 2016. Regulation of small RNAs and corresponding targets in Nod factor-induced Phaseolus vulgaris root hair cells. Int J Mol Sci . 176:887.
- ^ a b c d Jeffries, P.; Gianinazzi, S.; Perotto, S.; Turnau, K.; Barea, J. (2003). "The Contribution of arbuscular mycorrhizal fungi in sustainable maintenance of plant health and soil fertility". Biology and Fertility of Soils. 37 (1): 1–16. Bibcode:2003BioFS..37....1J. doi:10.1007/s00374-002-0546-5. S2CID 20792333.
- ^ Akhtar, Ovaid; Mishra, Rani; Kehri, Harbans Kaur (2019-03-01). "Arbuscular Mycorrhizal Association Contributes to Cr Accumulation and Tolerance in Plants Growing on Cr Contaminated Soils". Proceedings of the National Academy of Sciences, India Section B: Biological Sciences. 89 (1): 63–70. doi:10.1007/s40011-017-0914-4. ISSN 2250-1746. S2CID 46007322.
- ^ Fillion, M.; Brisson, J.; Guidi W.; Labrecque, M. (2011). "Increasing phosphorus removal in willow and poplar vegetation filters using arbuscular mycorrhizal fungi". Ecological Engineering. 37 (2): 199–205. Bibcode:2011EcEng..37..199F. doi:10.1016/j.ecoleng.2010.09.002.
- ^ Akhtar, Ovaid; Kehri, Harbans Kaur; Zoomi, Ifra (2020-09-15). "Arbuscular mycorrhiza and Aspergillus terreus inoculation along with compost amendment enhance the phytoremediation of Cr-rich technosol by Solanum lycopersicum under field conditions". Ecotoxicology and Environmental Safety. 201: 110869. Bibcode:2020EcoES.20110869A. doi:10.1016/j.ecoenv.2020.110869. ISSN 0147-6513. PMID 32585490. S2CID 220073862.
- ^ Giovannini, Luca; Palla, Michela; Agnolucci, Monica; Avio, Luciano; Sbrana, Cristiana; Turrini, Alessandra; Giovannetti, Manuela (2020). "Arbuscular Mycorrhizal Fungi and Associated Microbiota as Plant Biostimulants: Research Strategies for the Selection of the Best Performing Inocula". Agronomy. 10: 106. doi:10.3390/agronomy10010106. This article incorporates text from this source, which is available under the CC BY 4.0 license.
- ^ Rillig, M.; Ramsey, P.; Morris, S.; Paul, E. (2003). "Glomalin, an arbuscular-mycorrhizal fungal soil protein, responds to land-use change". Plant and Soil. 253 (2): 293–299. doi:10.1023/A:1024807820579. S2CID 11007821.
- ^ a b c Rillig, M. (2004). "Arbuscular mycorrhizae, glomalin and soil aggregation". Canadian Journal of Soil Science. 84 (4): 355–363. doi:10.4141/S04-003.
Notes
edit- Boswell, E. P.; R.T. Koide; D.L. Shumway; H.D. Addy. (1998). "Winter Wheat cover cropping, VA mycorrhizal fungi and maize growth and yield". Agriculture, Ecosystems and Environment. 67 (1): 55–65. Bibcode:1998AgEE...67...55B. doi:10.1016/S0167-8809(97)00094-7.
- Bücking H.; Shachar-Hill Y. (2005). "Phosphate uptake, transport and transfer by arbuscular mycorrhizal fungus is increased by carbohydrate availability". New Phytologist. 165 (3): 889–912. doi:10.1111/j.1469-8137.2004.01274.x. PMID 15720701.
- George E., K. Haussler, S.K. Kothari, X.L. Li and H. Marshner,1992 Contribution of Mycorrhizal Hyphae to Nutrient and Water Uptake of Plants. In Mycorrhizas in Ecosystems, ed., D.J. Read, D.H. Lewis, A.H. Fitter, I.J. Alexander. United Kingdom: C.A.B. International, pp. 42–47.
- Grant, C.; Bitman, S.; Montreal, M.; Plenchette, C.; Morel, C. (2005). "Soil and fertilizer phosphorus: effects on plant supply and mycorrhizal development". Canadian Journal of Plant Science. 85: 3–14. doi:10.4141/P03-182.
- Kosuta, S.; Chabaud, M.; Lougnon, G.; Gough, C.; Denarie, J.; Barker, D.; Bacard, G. (2003). "A Diffusible Factor from Arbuscular Mycorrhizal Fungi Induces Symbiosis-Specific MtENOD11 Expression in Roots of Medicago truncatula". Plant Physiology. 131 (3): 952–962. doi:10.1104/pp.011882. PMC 166861. PMID 12644648.
- Kabir, Z.; R.T. Koide (2000). "The effect of dandelion or a cover crop on mycorrhiza inoculum potential, soil aggregation and yield of maize". Agriculture, Ecosystems and Environment. 78 (2): 167–174. Bibcode:2000AgEE...78..167K. doi:10.1016/S0167-8809(99)00121-8.
- McGonigle, T.P.; M.H. Miller (1999). "Winter survival of extraradical hyphae and spores of arbuscular mycorrhizal fungi in the field". Applied Soil Ecology. 12 (1): 41–50. Bibcode:1999AppSE..12...41M. doi:10.1016/S0929-1393(98)00165-6.
- Miller, M.H.; McGonigle T.P.; Addy, H.D. (1995). "Functional ecology if vesicular arbuscular mycorrhizas as influenced by phosphate fertilization and tillage in an agricultural ecosystem". Critical Reviews in Biotechnology. 15 (3–4): 241–255. doi:10.3109/07388559509147411.
- Mozafar, A.; Anken, T.; Ruh, R.; Frossard, E. (2000). "Tillage intensity, Mycorrhizal and non mycorrhizal fungi and nutrient concentrations in maize, wheat and canola". Agronomy Journal. 92 (6): 1117–1124. Bibcode:2000AgrJ...92.1117M. doi:10.2134/agronj2000.9261117x.
- Sorensen, J.N.; J Larsen; I. Jakobsen (2005). "Mycorrhizae formation and nutrient concentration in leeks (Allium porrum) in relation to previous crop and cover crop management on high P soils". Plant and Soil. 273 (1–2): 101–114. Bibcode:2005PlSoi.273..101S. doi:10.1007/s11104-004-6960-8. S2CID 30777851.
- Thingstrup, I.; G. Rubaek; E. Sibbensen; I. Jakobsen (1999). "Flax (Linum usitatissimum L.) depends on arbuscular mycorrhizal fungi for growth and P uptake at intermediate but not high soil P levels in the field". Plant and Soil. 203: 37–46. doi:10.1023/A:1004362310788. S2CID 27345855.
- Timmer, L.; Leyden, R. (1980). "The relationship of mycorrhizal infection to phosphorus-induced copper deficiency in sour orange seedlings". New Phytologist. 85: 15–23. doi:10.1111/j.1469-8137.1980.tb04443.x. S2CID 85946706.
- Xie, Z.; Staehelin, C.; Vierheilig, H.; Weimken, A.; Jabbouri, S.; Broughton W.; Vogeli-Lange, R.; Thomas B. (1995). "Rhizobial Nodulation Factors Stimulate Mycorrhizal Colonization of Nodulating and Nonnodulating Soybeans". Plant Physiology. 108 (4): 1519–1525. doi:10.1104/pp.108.4.1519. PMC 157531. PMID 12228558.