DNA base flipping, or nucleotide flipping, is a mechanism in which a single nucleotide base, or nucleobase, is rotated outside the nucleic acid double helix.[1] This occurs when a nucleic acid-processing enzyme needs access to the base to perform work on it, such as its excision for replacement with another base during DNA repair. It was first observed in 1994 using X-ray crystallography in a methyltransferase enzyme catalyzing methylation of a cytosine base in DNA. Since then, it has been shown to be used by different enzymes in many biological processes such as DNA methylation, various DNA repair mechanisms, and DNA replication. It can also occur in RNA double helices [2] or in the DNA:RNA intermediates formed during RNA transcription.
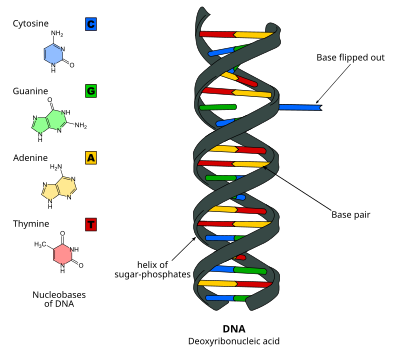
DNA base flipping occurs by breaking the hydrogen bonds between the bases and unstacking the base from its neighbors. This could occur through an active process, where an enzyme binds to the DNA and then facilitates rotation of the base, or a passive process, where the base rotates out spontaneously, and this state is recognized and bound by an enzyme. It can be detected using X-ray crystallography, NMR spectroscopy, fluorescence spectroscopy, or hybridization probes.
Discovery
editBase flipping was first observed in 1994 when researchers Klimasauskas, Kumar, Roberts, and Cheng used X-ray crystallography to view an intermediate step in the chemical reaction of a methyltransferase bound to DNA.[3] The methyltransferase they used was the C5-cytosine methyltransferase from Haemophilus haemolyticus (M. HhaI). This enzyme recognizes a specific sequence of the DNA (5'-GCGC-3') and methylates the first cytosine base of the sequence at its C5 location.[3] Upon crystallization of the M. HhaI-DNA complex, they saw the target cytosine base was rotated completely out of the double helix and was positioned in the active site of the M. HhaI. It was held in place by numerous interactions between the M. HhaI and DNA.[3]
The authors theorized that base flipping was a mechanism used by many other enzymes, such as helicases, recombination enzymes, RNA polymerases, DNA polymerases, and Type II topoisomerases.[3] Much research has been done in the years subsequent to this discovery and it has been found that base flipping is a mechanism used in many of the biological processes the authors suggest.[4][5][6]
Mechanism
editDNA nucleotides are held together with hydrogen bonds, which are relatively weak and can be easily broken. Base flipping occurs on a millisecond timescale[7] by breaking the hydrogen bonds between bases and unstacking the base from its neighbors.[8] The base is rotated out of the double helix by 180 degrees.,[9] typically via the major groove,[10] and into the active site of an enzyme. This opening leads to small conformational changes in the DNA backbone[11] which are quickly stabilized by the increased enzyme-DNA interactions.[12] Studies looking at the free-energy profiles of base flipping have shown that the free-energy barrier to flipping can be lowered by 17 kcal/mol for M.HhaI in the closed conformation.[10]
There are two mechanisms of DNA base flipping: active and passive.[13] In the active mechanism, an enzyme binds to the DNA and then actively rotates the base, while in the passive mechanism a damaged base rotates out spontaneously first, then is recognized and bound by the enzyme.[8] Research has demonstrated both mechanisms: uracil-DNA glycosylase follows the passive mechanism[8] and Tn10 transposase follows the active mechanism.[14]
Furthermore, studies have shown that DNA base flipping is used by many different enzymes in a variety biological processes such as DNA methylation, various DNA repair mechanisms, RNA transcription and DNA replication.[4][5][6]
Biological processes
editDNA modification and repair
editDNA can have mutations that cause a base in the DNA strand to be damaged. To ensure genetic integrity of the DNA, enzymes need to repair any damage. There are many types of DNA repair. Base excision repair utilizes base flipping to flip the damaged base out of the double helix[5] and into the specificity pocket of a glycosylase which hydrolyzes the glycosidic bond and removes the base.[15] DNA glycosylases interact with DNA, flipping bases to determine a mismatch. An example of base excision repair occurs when a cytosine base is deaminated and becomes a uracil base. This causes a U:G mispair which is detected by Uracil DNA glycosylase. The uracil base is flipped out into the glycosylase active pocket where it is removed from the DNA strand.[16] Base flipping is used to repair mutations such as 8-Oxoguanine (oxoG)[17] and thymine dimers created by UV radiation.[15][18]
Replication, transcription and recombination
editDNA replication and RNA transcription both make use of base flipping.[5] DNA polymerase is an enzyme that carries out replication. It can be thought of as a hand that grips the DNA single strand template.[15] As the template passes across the palm region of the polymerase, the template bases are flipped out of the helix and away from the dNTP binding site.[19] During transcription, RNA polymerase catalyzes RNA synthesis. During the initiation phase, two bases in the -10 element flip out from the helix and into two pockets in RNA polymerase. These new interactions stabilize the -10 element and promote the DNA strands to separate or melt.[15][20]
Base flipping occurs during latter stages of recombination.[21] RecA is a protein that promotes strand invasion[15] during homologous recombination. Base flipping has been proposed as the mechanism by which RecA can enable a single strand to recognize homology in duplex DNA.[22] Other studies indicate that it is also involved in V(D)J Recombination.[23]
DNA methylation
editDNA methylation is the process in which a methyl group is added to either a cytosine or adenine.[24] This process causes the activation or inactivation of gene expression, thereby resulting in gene regulation in eukaryotic cells. DNA methylation process is also known to be involved in certain types of cancer formation.[25][26][27] In order for this chemical modification to occur, it is necessary that the target base flips out of the DNA double helix to allow the methyltransferases to catalyze the reaction.[5]
Target recognition by restriction endonucleases
editRestriction endonucleases, also known as restriction enzymes are enzymes that cleave the sugar-phosphate backbone of the DNA at specific nucleotides sequences that are usually four to six nucleotides long.[28] Studies performed by Horton and colleagues have shown that the mechanism by which these enzymes cleave the DNA involves base flipping as well as bending the DNA and the expansion of the minor groove.[29] In 2006, Horton and colleagues, x-ray crystallography evidence was presented showing that the restriction endonuclease HinP1I utilizes base flipping in order to recognize its target sequence. This enzyme is known to cleave the DNA at the palindromic tetranucleotide sequence G↓CGC.
Experimental approaches for detection
editX-ray crystallography
editX-ray crystallography is a technique that measures the angles and intensities of crystalline atoms in order to determine the atomic and molecular structure of the crystal of interest. Crystallographers are then able to produce and three-dimensional picture where the positions of the atoms, chemical bonds as well as other important characteristics can be determined.[30] Klimasaukas and colleagues used this technique to observe the first base flipping phenomenon, in which their experimental procedure involved several steps:[3]
- Purification
- Crystallization
- Data Collection
- Structure determination and refinement
During purification, Haemophilus haemolyticus methyltransferase was overexpressed and purified using a high salt back-extraction step to selectively solubilize M.HhaI, followed by fast protein liquid chromatography (FPLC) as done previously by Kumar and colleagues.[31] Authors utilized a Mono-Q anion exchange column to remove the small quantity of proteinaceous materials and unwanted DNA prior to the crystallization step. Once M.HhaI was successfully purified, the sample was then grown using a method that mixes the solution containing the complex at a temperature of 16 °C and the hanging-drop vapor diffusion technique to obtain the crystals. Authors were then able to collect the x-ray data according to a technique used by Cheng and colleagues in 1993.[32] This technique involved the measurement of the diffraction intensities on a FAST detector, where the exposure times for 0.1° rotation were 5 or 10 seconds. For the structure determination and refinement, Klimasaukas and colleagues used the molecular replacement of the refined apo structure described by Cheng and colleagues in 1993[32] where the search models X-PLOR, MERLOT, and TRNSUM were used to solve the rotation and translation functions.[33][34] This part of the study involves the use of a variety of software and computer algorithms to solve the structures and characteristics of the crystal of interest.
NMR spectroscopy
editNMR spectroscopy is a technique that has been used over the years to study important dynamic aspects of base flipping. This technique allows researchers to determine the physical and chemical properties of atoms and other molecules by utilizing the magnetic properties of atomic nuclei.[35] In addition, NMR can provide a variety of information including structure, reaction states, chemical environment of the molecules, and dynamics.[36][37] During the DNA base flipping discovery experiment, researchers utilized NMR spectroscopy to investigate the enzyme-induced base flipping of HhaI methyltransferase. In order to accomplish this experiment, two 5-fluorocytosine residues were incorporated into the target and the reference position with the DNA substrate so the 19F chemical shift analysis could be performed. Once the 19F chemical shift analysis was evaluated, it was then concluded that the DNA complexes existed with multiple forms of the target 5-fluorocytosine along the base flipping pathway.[38]
Fluorescence spectroscopy
editFluorescence spectroscopy is a technique that is used to assay a sample using a fluorescent probe. DNA nucleotides themselves are not good candidates for this technique because they do not readily re-emit light upon light excitation.[39] A fluorescent marker is needed to detect base flipping. 2-Aminopurine is a base that is structurally similar to adenine, but is very fluorescent when flipped out from the DNA duplex.[40] It is commonly used to detect base flipping and has an excitation at 305‑320 nm and emission at 370 nm so that it well separated from the excitations of proteins and DNA. Other fluorescent probes used to study DNA base flipping are 6MAP (4‑amino‑6‑methyl‑7(8H)‑pteridone)[41] and Pyrrolo‑C (3-[β-D-2-ribofuranosyl]-6-methylpyrrolo[2,3-d]pyrimidin-2(3H)-one).[42][43] Time-resolved fluorescence spectroscopy is also employed to provide a more detailed picture of the extent of base flipping as well as the conformational dynamics occurring during base flipping.[44]
Hybridization probing
editHybridization probes can be used to detect base flipping. This technique uses a molecule that has a complementary sequence to the sequence you would like to detect such that it binds to a single-strand of the DNA or RNA. Several hybridization probes have been used to detect base flipping. Potassium permanganate is used to detect thymine residues that have been flipped out by cytosine-C5 and adenine-N6 methyltransferases.[45] Chloroacetaldehyde is used to detect cytosine residues flipped out by the HhaI DNA cytosine-5 methyltransferase (M. HhaI).[46]
See also
editReferences
edit- ^ Roberts, RJ; Cheng, X (1998). "Base flipping". Annual Review of Biochemistry. 67 (1): 181–198. doi:10.1146/annurev.biochem.67.1.181. PMID 9759487.
- ^ Reiter, NJ; Blad, H; Abildgaard, F; Butcher, SE (2004). "Dynamics in the U6 RNA intramolecular stem-loop: a base flipping conformational change". Biochemistry. 43 (43): 13739–47. doi:10.1021/bi048815y. PMID 15504036. S2CID 25391616.
- ^ a b c d e Klimasauskas, Saulius; Kumar, Sanjay; Roberts, Richard J.; Cheng, Xiaodong (January 1994). "Hhal methyltransferase flips its target base out of the DNA helix". Cell. 76 (2): 357–369. doi:10.1016/0092-8674(94)90342-5. PMID 8293469. S2CID 23161543.
- ^ a b Brown, Tom. "Nucleic Acids Book". ATDBio. Retrieved 26 February 2014.
- ^ a b c d e Huang, Niu; Nilesh K. Banavali; Alexander D. MacKerell (January 7, 2003). "Protein-facilitated base flipping in DNA by cytosine-5-methyltransferase". PNAS. 100 (1): 68–73. Bibcode:2003PNAS..100...68H. doi:10.1073/pnas.0135427100. PMC 140885. PMID 12506195.
- ^ a b Grubmüller, Helmut. "DNA Base Flipping". Archived from the original on 4 February 2017. Retrieved 26 February 2014.
- ^ Bouvier, Benjamin; Grubmüller, Helmut (August 2007). "A Molecular Dynamics Study of Slow Base Flipping in DNA using Conformational Flooding" (PDF). Biophysical Journal. 93 (3): 770–786. Bibcode:2007BpJ....93..770B. doi:10.1529/biophysj.106.091751. PMC 1913169. PMID 17496048. Archived from the original (PDF) on 2017-08-09. Retrieved 2014-03-15.
- ^ a b c Lariviere, L. (23 June 2004). "Structural Evidence of a Passive Base-flipping Mechanism for Glucosyltransferase". Journal of Biological Chemistry. 279 (33): 34715–34720. doi:10.1074/jbc.M404394200. PMID 15178685.
- ^ Grosjean, [edited by] Henri (2009). DNA and RNA modification enzymes : structure, mechanism, function and evolution. Austin, Tex.: Landes Bioscience. ISBN 978-1-58706-329-9. Archived from the original on 2014-04-07. Retrieved 2014-03-10.
{{cite book}}:|first=has generic name (help) - ^ a b Huang, N.; Banavali, N. K.; MacKerell, A. D. (27 December 2002). "Protein-facilitated base flipping in DNA by cytosine-5-methyltransferase". Proceedings of the National Academy of Sciences. 100 (1): 68–73. doi:10.1073/pnas.0135427100. PMC 140885. PMID 12506195.
- ^ Giudice, E. (1 March 2003). "Base pair opening within B-DNA: free energy pathways for GC and AT pairs from umbrella sampling simulations". Nucleic Acids Research. 31 (5): 1434–1443. doi:10.1093/nar/gkg239. PMC 149832. PMID 12595551.
- ^ Huang, N.; Banavali, N. K.; MacKerell, A. D. (27 December 2002). "Protein-facilitated base flipping in DNA by cytosine-5-methyltransferase". Proceedings of the National Academy of Sciences. 100 (1): 68–73. doi:10.1073/pnas.0135427100. PMC 140885. PMID 12506195.
- ^ O'Neil, Lauren. Base Flipping in DNA: Detection, Structures and Energetics, A Dissertation. ISBN 9780549590743. Retrieved 15 March 2014.[permanent dead link]
- ^ Bischerour, Julien; Chalmers, Ronald; Bielinsky, Anja-Katrin (10 July 2009). "Base Flipping in Tn10 Transposition: An Active Flip and Capture Mechanism". PLOS ONE. 4 (7): e6201. Bibcode:2009PLoSO...4.6201B. doi:10.1371/journal.pone.0006201. PMC 2705183. PMID 19593448.
- ^ a b c d e University, James D. Watson, Cold Spring Harbor Laboratory, Tania A. Baker, Massachusetts Institute of Technology, Alexander Gann, Cold Spring Harbor Laboratory, Michael Levine, University of California, Berkeley, Richard Losik, Harvard (2014). Molecular biology of the gene (Seventh ed.). Boston: Pearson/CSH Press. ISBN 978-0-321-76243-6.
{{cite book}}: CS1 maint: multiple names: authors list (link) - ^ Krokan, Hans E; Drabløs, Finn; Slupphaug, Geir (16 December 2002). "Uracil in DNA – occurrence, consequences and repair". Oncogene. 21 (58): 8935–8948. doi:10.1038/sj.onc.1205996. PMID 12483510.
- ^ Banerjee, Anirban; Yang, Wei; Karplus, Martin; Verdine, Gregory L. (31 March 2005). "Structure of a repair enzyme interrogating undamaged DNA elucidates recognition of damaged DNA". Nature. 434 (7033): 612–618. Bibcode:2005Natur.434..612B. doi:10.1038/nature03458. PMID 15800616. S2CID 4426014.
- ^ Fuxreiter, Monika; Luo, Ning; Jedlovszky, Pál; Simon, István; Osman, Roman (November 2002). "Role of Base Flipping in Specific Recognition of Damaged DNA by Repair Enzymes". Journal of Molecular Biology. 323 (5): 823–834. doi:10.1016/S0022-2836(02)00999-3. PMID 12417196.
- ^ Patel, Premal H.; Suzuki, Motoshi; Adman, Elinor; Shinkai, Akeo; Loeb, Lawrence A. (May 2001). "Prokaryotic DNA polymerase I: evolution, structure, and "base flipping" mechanism for nucleotide selection". Journal of Molecular Biology. 308 (5): 823–837. doi:10.1006/jmbi.2001.4619. PMID 11352575. S2CID 16277925.
- ^ Lim, H. M.; Lee, H. J.; Roy, S.; Adhya, S. (4 December 2001). "A "master" in base unpairing during isomerization of a promoter upon RNA polymerase binding". Proceedings of the National Academy of Sciences. 98 (26): 14849–14852. Bibcode:2001PNAS...9814849L. doi:10.1073/pnas.261517398. PMC 64947. PMID 11734629.
- ^ Voloshin, Oleg N.; Camerini-Otero, R.Daniel (September 2004). "Synaptic Complex Revisited". Molecular Cell. 15 (6): 846–847. doi:10.1016/j.molcel.2004.09.010. PMID 15383274.
- ^ Folta-Stogniew, E; O'Malley, S; Gupta, R; Anderson, KS; Radding, CM (Sep 24, 2004). "Exchange of DNA base pairs that coincides with recognition of homology promoted by E. coli RecA protein". Molecular Cell. 15 (6): 965–75. doi:10.1016/j.molcel.2004.08.017. PMID 15383285.
- ^ Bischerour, J.; Lu, C.; Roth, D. B.; Chalmers, R. (31 August 2009). "Base Flipping in V(D)J Recombination: Insights into the Mechanism of Hairpin Formation, the 12/23 Rule, and the Coordination of Double-Strand Breaks". Molecular and Cellular Biology. 29 (21): 5889–5899. doi:10.1128/MCB.00187-09. PMC 2772739. PMID 19720743.
- ^ Klose, Robert J.; Adrian P. Bird (2006). "Genomic DNA methylation: the mark and its mediators". Trends in Biochemical Sciences. 31 (2): 89–97. doi:10.1016/j.tibs.2005.12.008. ISSN 0968-0004. PMID 16403636.
- ^ Nakao, M (2001). "Epigenetics: Interaction of DNA Methylation and Chromatin". Gene. 278 (1–2): 25–31. doi:10.1016/s0378-1119(01)00721-1. PMID 11707319.
- ^ Plass, C; Soloway, PD (2002). "DNA Methylation, Imprinting and Cancer". Eur J Hum Genet. 10 (1): 6–16. doi:10.1038/sj.ejhg.5200768. PMID 11896451.
- ^ Esteller, M; Herman, JG (2002). "Cancer as an Epigenetic Disease: DNA Methylation and Chromatin Alterations in Human Tumours". J Pathol. 196 (1): 1–7. doi:10.1002/path.1024. PMID 11748635. S2CID 35380651.
- ^ "Biology and Activity of Restriction Endonucleases". Archived from the original on 2014-04-18. Retrieved 2014-04-03.
- ^ Horton, John R.; Zhang, Xing; Maunus, Robert; Yang, Zhe; Wilson, Geoffrey; Roberts, Richard; Cheng, Xiaodong (2006). "DNA Nicking by HinP1I Endonuclease: Bending, Base Flipping and Minor Groove Expansion". Nucleic Acids Research. 34 (3): 939–948. doi:10.1093/nar/gkj484. PMC 1363774. PMID 16473850.
- ^ X-ray crystallography
- ^ Kumar, S; Cheng, X; Pflugrath, JW; Roberts, RJ (1992). "Purification, Crystallization, and Preliminary X-ray Diffraction Analysis of an M.HhaI-AdoMet Complex". Biochemistry. 31 (36): 8648–8653. doi:10.1021/bi00151a035. PMID 1390649.
- ^ a b Cheng, X; Kumar, S; Posfai, J; Pflugrath, JW; Roberts, RJ (1993). "Crystal Structure of the HhaI DNA Methyltransferase Complexed with S-adenosyl-L-methionine". Cell. 74 (2): 299–307. doi:10.1016/0092-8674(93)90421-l. PMID 8343957. S2CID 54238106.
- ^ Brunger A.T. (1992)"X-PLOR, Version 3.1 : A system for x-ray crystallography and NMR"(New Haven, Connecticut: Yale University Press)
- ^ Fitzgerald, P.M.D. (1988). "MERLOT, an integral package of computer programs for the determination of crystal structure by molecular replacement". J. Appl. Crystallogr. 21 (3): 273–288. doi:10.1107/s0021889887012299.
- ^ NMR spectroscopy
- ^ Gueron, M., and J. L. Leroy. 1995. Studies of basepair kinetics by NMR measurement of proton exchange. In Nuclear Magnetic Resonance And Nucleic Acids. Academic Press, San Diego, CA.
- ^ Leijon, M.; Graslund, A. (1992). "Effects of sequence and length on imino proton-exchange and basepair opening kinetics in DNA oligonucleotide duplexes". Nucleic Acids Res. 20 (20): 5339–5343. doi:10.1093/nar/20.20.5339. PMC 334339. PMID 1331987.
- ^ Klimasaukas, Salius, and Zita Liutkeviciute. "Experimental Approaches to Study DNA Base Flipping." DNA and RNA Modification Enzymes: Structure, Mechanism, Function and Evolution. Landes Bioscience, 2009. 37-50. Web. 16 Mar. 2014. <https://www.landesbioscience.com/pdf/04GrosjeanKlimasauskas.pdf Archived 2014-04-07 at the Wayback Machine>.
- ^ Grosjean, [edited by] Henri (2009). DNA and RNA modification enzymes : structure, mechanism, function and evolution (PDF). Austin, Tex.: Landes Bioscience. p. 43. ISBN 978-1-58706-329-9.
{{cite book}}:|first=has generic name (help) - ^ Holz, B (15 February 1998). "2-Aminopurine as a fluorescent probe for DNA base flipping by methyltransferases". Nucleic Acids Research. 26 (4): 1076–1083. doi:10.1093/nar/26.4.1076. PMC 147370. PMID 9461471.
- ^ Yang, K; Matsika, S; Stanley, RJ (Sep 6, 2007). "6MAP, a fluorescent adenine analogue, is a probe of base flipping by DNA photolyase". The Journal of Physical Chemistry B. 111 (35): 10615–25. doi:10.1021/jp071035p. PMID 17696385. S2CID 4998287.
- ^ Yang, K; Stanley, RJ (May–Jun 2008). "The extent of DNA deformation in DNA photolyase-substrate complexes: a solution state fluorescence study". Photochemistry and Photobiology. 84 (3): 741–9. doi:10.1111/j.1751-1097.2007.00251.x. PMID 18086248. S2CID 44506405.
- ^ Berry, David A.; Jung, Kee-Yong; Wise, Dean S.; Sercel, Anthony D.; Pearson, William H.; Mackie, Hugh; Randolph, John B.; Somers, Robert L. (March 2004). "Pyrrolo-dC and pyrrolo-C: fluorescent analogs of cytidine and 2′-deoxycytidine for the study of oligonucleotides". Tetrahedron Letters. 45 (11): 2457–2461. doi:10.1016/j.tetlet.2004.01.108.
- ^ Neely, R. K.; Tamulaitis, G.; Chen, K.; Kubala, M.; Siksnys, V.; Jones, A. C. (8 September 2009). "Time-resolved fluorescence studies of nucleotide flipping by restriction enzymes". Nucleic Acids Research. 37 (20): 6859–6870. doi:10.1093/nar/gkp688. PMC 2777440. PMID 19740769.
- ^ Serva, S (1 August 1998). "Chemical display of thymine residues flipped out by DNA methyltransferases". Nucleic Acids Research. 26 (15): 3473–3479. doi:10.1093/nar/26.15.3473. PMC 147733. PMID 9671807.
- ^ Daujotyte, D.; Liutkeviciute, Z.; Tamulaitis, G.; Klimasauskas, S. (15 April 2008). "Chemical mapping of cytosines enzymatically flipped out of the DNA helix". Nucleic Acids Research. 36 (10): e57. doi:10.1093/nar/gkn200. PMC 2425465. PMID 18450817.