Methyltransferases are a large group of enzymes that all methylate their substrates but can be split into several subclasses based on their structural features. The most common class of methyltransferases is class I, all of which contain a Rossmann fold for binding S-Adenosyl methionine (SAM). Class II methyltransferases contain a SET domain, which are exemplified by SET domain histone methyltransferases, and class III methyltransferases, which are membrane associated.[1] Methyltransferases can also be grouped as different types utilizing different substrates in methyl transfer reactions. These types include protein methyltransferases, DNA/RNA methyltransferases, natural product methyltransferases, and non-SAM dependent methyltransferases. SAM is the classical methyl donor for methyltransferases, however, examples of other methyl donors are seen in nature. The general mechanism for methyl transfer is a SN2-like nucleophilic attack where the methionine sulfur serves as the leaving group and the methyl group attached to it acts as the electrophile that transfers the methyl group to the enzyme substrate. SAM is converted to S-Adenosyl homocysteine (SAH) during this process. The breaking of the SAM-methyl bond and the formation of the substrate-methyl bond happen nearly simultaneously. These enzymatic reactions are found in many pathways and are implicated in genetic diseases, cancer, and metabolic diseases. Another type of methyl transfer is the radical S-Adenosyl methionine (SAM) which is the methylation of unactivated carbon atoms in primary metabolites, proteins, lipids, and RNA.
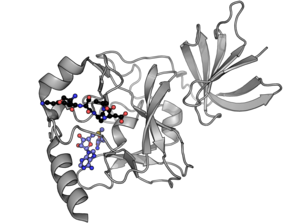
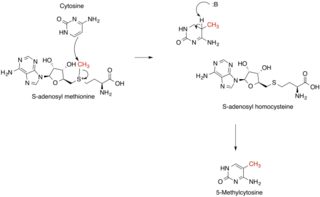
Function
editGenetics
editMethylation, as well as other epigenetic modifications, affects transcription, gene stability, and parental imprinting.[2] It directly impacts chromatin structure and can modulate gene transcription, or even completely silence or activate genes, without mutation to the gene itself. Though the mechanisms of this genetic control are complex, hypo- and hypermethylation of DNA is implicated in many diseases.
Protein regulation
editMethylation of proteins has a regulatory role in protein–protein interactions, protein–DNA interactions, and protein activation.
Examples: RCC1, an important mitotic protein, is methylated so that it can interact with centromeres of chromosomes. This is an example of regulation of protein-protein interaction, as methylation regulates the attachment of RCC1 to histone proteins H2A and H2B. The RCC1-chromatin interaction is also an example of a protein-DNA interaction, as another domain of RCC1 interacts directly with DNA when this protein is methylated. When RCC1 is not methylated, dividing cells have multiple spindle poles and usually cannot survive.
p53 methylated on lysine to regulate its activation and interaction with other proteins in the DNA damage response. This is an example of regulation of protein-protein interactions and protein activation. p53 is a known tumor suppressor that activates DNA repair pathways, initiates apoptosis, and pauses the cell cycle. Overall, it responds to mutations in DNA, signaling to the cell to fix them or to initiate cell death so that these mutations cannot contribute to cancer.
NF-κB (a protein involved in inflammation) is a known methylation target of the methyltransferase SETD6, which turns off NF-κB signaling by inhibiting of one of its subunits, RelA. This reduces the transcriptional activation and inflammatory response, making methylation of NF-κB a regulatory process by which cell signaling through this pathway is reduced.[3]
Natural product methyltransferases provide a variety of inputs into metabolic pathways, including the availability of cofactors, signalling molecules, and metabolites. This regulates various cellular pathways by controlling protein activity.
Types
editHistone methyltransferases
editHistone methyltransferases are critical for genetic regulation at the epigenetic level. They modify mainly lysine on the ε-nitrogen and the arginine guanidinium group on histone tails. Lysine methyltransferases and Arginine methyltransferases are unique classes of enzymes, but both bind SAM as a methyl donor for their histone substrates. Lysine amino acids can be modified with one, two, or three methyl groups, while Arginine amino acids can be modified with one or two methyl groups. This increases the strength of the positive charge and residue hydrophobicity, allowing other proteins to recognize methyl marks. The effect of this modification depends on the location of the modification on the histone tail and the other histone modifications around it. The location of the modifications can be partially determined by DNA sequence, as well as small non-coding RNAs and the methylation of the DNA itself. Most commonly, it is histone H3 or H4 that is methylated in vertebrates. Either increased or decreased transcription of genes around the modification can occur. Increased transcription is a result of decreased chromatin condensation, while decreased transcription results from increased chromatin condensation.[4] Methyl marks on the histones contribute to these changes by serving as sites for recruitment of other proteins that can further modify chromatin.[5]
N-terminal methyltransferases
editN-alpha methyltransferases transfer a methyl group from SAM to the N-terminal nitrogen on protein targets. The N-terminal methionine is first cleaved by another enzyme and the X-Proline-Lysine consensus sequence is recognized by the methyltransferase. For all known substrates, the X amino acid is Alanine, Serine, or Proline. This reaction yields a methylated protein and SAH. Known targets of these methyltransferases in humans include RCC-1 (a regulator of nuclear transport proteins) and Retinoblastoma protein (a tumor suppressor protein that inhibits excessive cell division). RCC-1 methylation is especially important in mitosis as it coordinates the localization of some nuclear proteins in the absence of the nuclear envelope. When RCC-1 is not methylated, cell division is abnormal following the formation of extra spindle poles.[6] The function of Retinoblastoma protein N-terminal methylation is not known.
DNA/RNA methyltransferases
editDNA methylation, a key component of genetic regulation, occurs primarily at the 5-carbon of the base cytosine, forming 5’methylcytosine (see left).[7] Methylation is an epigenetic modification catalyzed by DNA methyltransferase enzymes, including DNMT1, DNMT2 (renamed TRDMT1 to reflect its function methylating tRNA, not DNA), and DNMT3. These enzymes use S-adenosylmethionine as a methyl donor and contain several highly conserved structural features between the three forms; these include the S-adenosylmethionine binding site, a vicinal proline-cysteine pair which forms a thiolate anion important for the reaction mechanism, and the cytosine substrate binding pocket. Many features of DNA methyltransferases are highly conserved throughout many classes of life, from bacteria to mammals. In addition to controlling the expression of certain genes, there are a variety of protein complexes, many with implications for human health, which only bind to methylated DNA recognition sites. Many of the early DNA methyltransferases have been thought to be derived from RNA methyltransferases that were supposed to be active in the RNA world to protect many species of primitive RNA.[8] RNA methylation has been observed in different types of RNA species viz.mRNA, rRNA, tRNA, snoRNA, snRNA, miRNA, tmRNA as well as viral RNA species. Specific RNA methyltransferases are employed by cells to mark these on the RNA species according to the need and environment prevailing around the cells, which form a part of field called molecular epigenetics. 2'-O-methylation, m6A methylation, m1G methylation as well as m5C are most commonly methylation marks observed in different types of RNA.
6A is an enzyme that catalyzes chemical reaction as following:[9]
S-adenosyl-L-methionine + DNA adenine S-adenosyl-L-homocysteine + DNA 6-methylaminopurine
m6A was primarily found in prokaryotes until 2015 when it was also identified in some eukaryotes. m6A methyltransferases methylate the amino group in DNA at C-6 position specifically to prevent the host system to digest own genome through restriction enzymes.[10]
m5C plays a role to regulate gene transcription. m5C transferases are the enzymes that produce C5-methylcytosine in DNA at C-5 position of cytosine and are found in most plants and some eukaryotes.[11]
Natural product methyltransferases
editNatural product methyltransferases (NPMTs) are a diverse group of enzymes that add methyl groups to naturally-produced small molecules. Like many methyltransferases, SAM is utilized as a methyl donor and SAH is produced. Methyl groups are added to S, N, O, or C atoms, and are classified by which of these atoms are modified, with O-methyltransferases representing the largest class. The methylated products of these reactions serve a variety of functions, including co-factors, pigments, signalling compounds, and metabolites. NPMTs can serve a regulatory role by modifying the reactivity and availability of these compounds. These enzymes are not highly conserved across different species, as they serve a more specific function in providing small molecules for specialized pathways in species or smaller groups of species. Reflective of this diversity is the variety of catalytic strategies, including general acid-base catalysis, metal-based catalysis, and proximity and desolvation effects not requiring catalytic amino acids. NPMTs are the most functionally diverse class of methyltransferases.[12]
Important examples of this enzyme class in humans include phenylethanolamine N-methyltransferase (PNMT), which converts norepinephrine to epinephrine,[14] and histamine N-methyltransferase (HNMT), which methylates histamine in the process of histamine metabolism.[15] Catechol-O-methyltransferase (COMT) degrades a class of molecules known as catecholamines that includes dopamine, epinephrine, and norepenepherine.[16]
Non-SAM dependent methyltransferases
editMethanol, methyl tetrahydrofolate, mono-, di-, and trimethylamine, methanethiol, methyltetrahydromethanopterin, and chloromethane are all methyl donors found in biology as methyl group donors, typically in enzymatic reactions using the cofactor vitamin B12.[17] These substrates contribute to methyl transfer pathways including methionine biosynthesis, methanogenesis, and acetogenesis.
Radical SAM methyltransferases
editBased on different protein structures and mechanisms of catalysis, there are 3 different types of radical SAM (RS) methylases: Class A, B, and C. Class A RS methylases are the best characterized of the 4 enzymes and are related to both RlmN and Cfr. RlmN is ubiquitous in bacteria which enhances translational fidelity and RlmN catalyzes methylation of C2 of adenosine 2503 (A2503) in 23 S rRNA and C2 of adenosine (A37). Cfr, on the other hand, catalyzes methylation of C8 of A2503 as well and it also catalyzes C2 methylation.[18] Class B is currently the largest class of radical SAM methylases which can methylate both sp2-hybridized and sp3-hybridized carbon atoms in different sets of substrates unlike Class A which only catalyzes sp2-hybridized carbon atoms. The main difference that distinguishes Class B from others is the additional N-terminal cobalamin-binding domain that binds to the RS domain.[19] Class C methylase has homologous sequence with the RS enzyme, coproporphyrinogen III oxidase (HemN), which also catalyzes the methylation of sp2-hybridized carbon centers yet it lacks the 2 cysteines required for methylation in mechanism of Class A.[18]
Clinical significance
editAs with any biological process which regulates gene expression and/or function, anomalous DNA methylation is associated with genetic disorders such as ICF, Rett syndrome, and Fragile X syndrome.[2] Cancer cells typically exhibit less DNA methylation activity in general, though often hypermethylation at sites which are unmethylated in normal cells; this overmethylation often functions as a way to inactivate tumor-suppressor genes. Inhibition of overall DNA methyltransferase activity has been proposed as a treatment option, but DNMT inhibitors, analogs of their cytosine substrates, have been found to be highly toxic due to their similarity to cytosine (see right); this similarity to the nucleotide causes the inhibitor to be incorporated into DNA translation, causing non-functioning DNA to be synthesized.
A methylase which alters the ribosomal RNA binding site of the antibiotic linezolid causes cross-resistance to other antibiotics that act on the ribosomal RNA. Plasmid vectors capable of transmitting this gene are a cause of potentially dangerous cross resistance.[20]
Examples of methyltransferase enzymes relevant to disease:
- thiopurine methyltransferase: defects in this gene causes toxic accumulation of thiopurine compounds, drugs used in chemotherapy and immunosuppressant therapy
- methionine synthase: pernicious anemia, caused by Vitamin B12 deficiency, is caused by a lack of cofactor for the methionine synthase enzyme
Applications in drug discovery and development
editRecent work has revealed the methyltransferases involved in methylation of naturally occurring anticancer agents to use S-Adenosyl methionine (SAM) analogs that carry alternative alkyl groups as a replacement for methyl. The development of the facile chemoenzymatic platform to generate and utilize differentially alkylated SAM analogs in the context of drug discovery and drug development is known as alkylrandomization.[21]
Applications in cancer treatment
editIn human cells, it was found that m5C was associated with abnormal tumor cells in cancer.[22] The role and potential application of m5C includes to balance the impaired DNA in cancer both hypermethylation and hypomethylation. An epigenetic repair of DNA can be applied by changing the m5C amount in both types of cancer cells (hypermethylation/ hypomethylation) and as well as the environment of the cancers to reach an equivalent point to inhibit tumor cells.[23]
Examples
editExamples include:
References
edit- ^ Katz, J. E.; Dlakic, M; Clarke S (18 July 2003). "Automated identification of putative methyltransferases from genomic open reading frames". Molecular & Cellular Proteomics. 2 (8): 525–40. doi:10.1074/mcp.M300037-MCP200. PMID 12872006.
- ^ a b Siedlecki, P; Zielenkiewicz, P (2006). "Mammalian DNA methyltransferases". Acta Biochimica Polonica. 53 (2): 245–56. doi:10.18388/abp.2006_3337. PMID 16582985.
- ^ Levy, Dan; et al. (5 December 2010). "Lysine methylation of the NF-κB subunit RelA by SETD6 couples activity of the histone methyltransferase GLP at chromatin to tonic repression of NF-κB signaling". Nature Immunology. 12 (1): 29–36. doi:10.1038/ni.1968. PMC 3074206. PMID 21131967.
- ^ Turner, Bryan M. (2001). Chromatin and gene regulation : mechanisms in epigenetics. Malden, MA: Blackwell Science. ISBN 978-0865427433.
- ^ Greer, Eric L.; Shi, Yang (3 April 2012). "Histone methylation: a dynamic mark in health, disease and inheritance". Nature Reviews Genetics. 13 (5): 343–357. doi:10.1038/nrg3173. PMC 4073795. PMID 22473383.
- ^ Clarke, Paul (May 2007). "Anchoring RCC1 by the tail". Nature Cell Biology. 9 (5): 485–487. doi:10.1038/ncb0507-485. PMID 17473856. S2CID 711645.
- ^ Lan, J; Hua, S; He, X; Zhang, Y (2010). "DNA methyltransferases and methyl-binding proteins of mammals". Acta Biochimica et Biophysica Sinica. 42 (4): 243–52. doi:10.1093/abbs/gmq015. PMID 20383462.
- ^ Rana, Ajay K.; Ankri, Serge (2016-01-01). "Reviving the RNA World: An Insight into the Appearance of RNA Methyltransferases". Frontiers in Genetics. 7: 99. doi:10.3389/fgene.2016.00099. PMC 4893491. PMID 27375676.
- ^ Kessler, Christoph; Manta, Vicentiu (1990-01-01). "Specificity of restriction endonucleases and DNA modification methyltransferases — a review (edition 3)". Gene. 92 (1): 1–240. doi:10.1016/0378-1119(90)90486-B. ISSN 0378-1119. PMID 2172084.
- ^ Narva, Kenneth E.; Van Etten, James L.; Slatko, Barton E.; Benner, Jack S. (1988-12-25). "The amino acid sequence of the eukaryotic DNA [N6-adenine]methyltransferase M·CviBIII, has regions of similarity with the prokaryotic isoschizomer M · TaqI and other DNA [N6-adenine] methyltransferases". Gene. 74 (1): 253–259. doi:10.1016/0378-1119(88)90298-3. ISSN 0378-1119. PMID 3248728.
- ^ Posfai, Janos; Bhagwat, Ashok S.; Roberts, Richard J. (1988-12-25). "Sequence motifs specific for cytosine methyltransferases". Gene. 74 (1): 261–265. doi:10.1016/0378-1119(88)90299-5. ISSN 0378-1119. PMID 3248729.
- ^ Liscombe, David K.; Louie, Gordon V.; Noel, Joseph P. (2012). "Architectures, mechanisms and molecular evolution of natural product methyltransferases". Natural Product Reports. 29 (10): 1238–50. doi:10.1039/c2np20029e. PMID 22850796.
- ^ Ashihara, Hiroshi; Yokota, Takao; Crozier, Alan (2013). Biosynthesis and catabolism of purine alkaloids. Advances in Botanical Research. Vol. 68. pp. 111–138. doi:10.1016/B978-0-12-408061-4.00004-3. ISBN 9780124080614.
- ^ "PNMT phenylethanolamine N-methyltransferase". NCBI Genetic Testing Registry. Retrieved 18 February 2014.
- ^ "HNMT histamine N-methyltransferase". NCBI Genetic Testing Registry. Retrieved 18 February 2014.
- ^ "COMT catechol-O-methyltransferase". NCBI Genetic Testing Registry. Retrieved 18 February 2014.
- ^ Ragsdale, S.W. "Catalysis of methyl group transfers involving tetrahydrofolate and B12" Vitamins and Hormones, 2008.
- ^ a b Bauerle, Matthew R.; Schwalm, Erica L.; Booker, Squire J. (2015-02-13). "Mechanistic Diversity of Radical S-Adenosylmethionine (SAM)-dependent Methylation". The Journal of Biological Chemistry. 290 (7): 3995–4002. doi:10.1074/jbc.R114.607044. ISSN 0021-9258. PMC 4326810. PMID 25477520.
- ^ Sofia, H. J.; Chen, G.; Hetzler, B. G.; Reyes-Spindola, J. F.; Miller, N. E. (2001-03-01). "Radical SAM, a novel protein superfamily linking unresolved steps in familiar biosynthetic pathways with radical mechanisms: functional characterization using new analysis and information visualization methods". Nucleic Acids Research. 29 (5): 1097–1106. doi:10.1093/nar/29.5.1097. ISSN 1362-4962. PMC 29726. PMID 11222759.
- ^ Morales G, Picazo JJ, Baos E, Candel FJ, Arribi A, Peláez B, Andrade R, de la Torre MA, Fereres J, Sánchez-García M (March 2010). "Resistance to linezolid is mediated by the cfr gene in the first report of an outbreak of linezolid-resistant Staphylococcus aureus". Clin. Infect. Dis. 50 (6): 821–5. doi:10.1086/650574. PMID 20144045.
- ^ Singh, S; Zhang, J; Huber, TD; Sunkara, M; Hurley, K; Goff, RD; Wang, G; Zhang, W; Liu, C; Rohr, J; Van Lanen, SG; Morris, AJ; Thorson, JS (7 April 2014). "Facile chemoenzymatic strategies for the synthesis and utilization of S-adenosyl-(L)-methionine analogues". Angewandte Chemie International Edition in English. 53 (15): 3965–9. doi:10.1002/anie.201308272. PMC 4076696. PMID 24616228.
- ^ Jones, Peter A. (1996-06-01). "DNA Methylation Errors and Cancer". Cancer Research. 56 (11): 2463–2467. ISSN 0008-5472. PMID 8653676.
- ^ D, Hanahan; Ra, Weinberg (2011-03-04). "Hallmarks of Cancer: The Next Generation". Cell. 144 (5): 646–74. doi:10.1016/j.cell.2011.02.013. PMID 21376230.
Further reading
edit- Methyltransferases at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- 3-D Structure of DNA Methyltransferase
- A novel methyltransferase : the 7SK snRNA Methylphosphate Capping Enzyme as seen on Flintbox
- "The Role of Methylation in Gene Expression" on Nature Scitable
- "Nutrition and Depression: Nutrition, Methylation, and Depression" on Psychology Today
- "DNA Methylation - What is DNA Methylation?" from News-Medical.net
- "Histone Lysine Methylation" Genetic pathways involving Histone Methyltransferases from Cell Signaling Technology