In molecular biology, the TATA box (also called the Goldberg–Hogness box)[1] is a sequence of DNA found in the core promoter region of genes in archaea and eukaryotes.[2] The bacterial homolog of the TATA box is called the Pribnow box which has a shorter consensus sequence.
The TATA box is considered a non-coding DNA sequence (also known as a cis-regulatory element). It was termed the "TATA box" as it contains a consensus sequence characterized by repeating T and A base pairs.[3] How the term "box" originated is unclear. In the 1980s, while investigating nucleotide sequences in mouse genome loci, the Hogness box sequence was found and "boxed in" at the -31 position.[4] When consensus nucleotides and alternative ones were compared, homologous regions were "boxed" by the researchers.[4] The boxing in of sequences sheds light on the origin of the term "box".
The TATA box was first identified in 1978[1] as a component of eukaryotic promoters. Transcription is initiated at the TATA box in TATA-containing genes. The TATA box is the binding site of the TATA-binding protein (TBP) and other transcription factors in some eukaryotic genes. Gene transcription by RNA polymerase II depends on the regulation of the core promoter by long-range regulatory elements such as enhancers and silencers.[5] Without proper regulation of transcription, eukaryotic organisms would not be able to properly respond to their environment.
Based on the sequence and mechanism of TATA box initiation, mutations such as insertions, deletions, and point mutations to this consensus sequence can result in phenotypic changes. These phenotypic changes can then turn into a disease phenotype. Some diseases associated with mutations in the TATA box include gastric cancer, spinocerebellar ataxia, Huntington's disease, blindness, β-thalassemia, immunosuppression, Gilbert's syndrome, and HIV-1. The TATA-binding protein (TBP) could also be targeted by viruses as a means of viral transcription.[6]
History
editDiscovery
editThe TATA box was the first eukaryotic core promoter motif to be identified in 1978 by American biochemist David Hogness[1] while he and his graduate student, Michael Goldberg were on sabbatical at the University of Basel in Switzerland.[7] They first discovered the TATA sequence while analyzing 5' DNA promoter sequences in Drosophila, mammalian, and viral genes.[8][2] The TATA box was found in protein coding genes transcribed by RNA polymerase II.[2]
Evolutionary history
editMost research on the TATA box has been conducted on yeast, human, and Drosophila genomes, however, similar elements have been found in archaea and ancient eukaryotes.[2] In archaea species, the promoter contains an 8 bp AT-rich sequence located ~24 bp upstream of the transcription start site. This sequence was originally called Box A, which is now known to be the sequence that interacts with the homologue of the archaeal TATA-binding protein (TBP). Also, even though some studies have uncovered several similarities, there are others that have detected notable differences between archaeal and eukaryotic TBP. The archaea protein exhibits a greater symmetry in its primary sequence and in the distribution of electrostatic charge, which is important because the higher symmetry lowers the protein's ability to bind the TATA box in a polar manner.[2]
Even though the TATA box is present in many eukaryotic promoters, it is not contained in the majority of promoters. One study found less than 30% of 1031 potential promoter regions contain a putative TATA box motif in humans.[9] In Drosophila, less than 40% of 205 core promoters contain a TATA box.[8] When there is an absence of the TATA box and TBP is not present, the downstream promoter element (DPE) in cooperation with the initiator element (Inr) bind to the transcription factor II D (TFIID), initiating transcription in TATA-less promoters. The DPE has been identified in three Drosophila TATA-less promoters and in the TATA-less human IRF-1 promoter.[10]
Features
editLocation
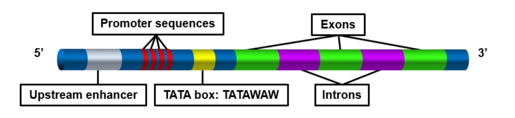
editPromoter sequences vary between bacteria and eukaryotes. In eukaryotes, the TATA box is located 25 base pairs upstream of the start site that Rpb4/Rbp7 use to initiate transcription. In metazoans, the TATA box is located 30 base pairs upstream of the transcription start site.[5] While in yeast, S. cerevisiae, the TATA box has a variable position which can range from 40 to 100 bp upstream of the start site. The TATA box is also found in 40% of the core promoters of genes that code for the actin cytoskeleton and contractile apparatus in cells.[5]
The type of core promoter affects the level of transcription and expression of a gene. TATA-binding protein (TBP) can be recruited in two ways, by SAGA, a cofactor for RNA polymerase II, or by TFIID.[11] When promoters use the SAGA/TATA box complex to recruit RNA polymerase II, they are more highly regulated and display higher expression levels than promoters using the TFIID/TBP mode of recruitment.[11]
Analogous sequences
editIn bacteria, promoter regions may contain a Pribnow box, which serves an analogous purpose to the eukaryotic TATA box. The Pribnow box has a 6 bp region centered around the -10 position and an 8-12 bp sequence around the -35 region that are both conserved.[10]
A CAAT box (also CAT box) is a region of nucleotides with the following consensus sequence: 5’ GGCCAATCT 3’. The CAAT box is located about 75-80 bases upstream of the transcription initiation site and about 150 bases upstream of the TATA box. It binds transcription factors (CAAT TF or CTFs) and thereby stabilizes the nearby preinitiation complex for easier binding of RNA polymerases. CAAT boxes are rarely found in genes that express proteins ubiquitous in all cell types.[10]
Structure
editSequence and prevalence
editThe TATA box is a component of the eukaryotic core promoter and generally contains the consensus sequence 5'-TATA(A/T)A(A/T)-3'.[3] In yeast, for example, one study found that various Saccharomyces genomes had the consensus sequence 5'-TATA(A/T)A(A/T)(A/G)-3', yet only about 20% of yeast genes even contained the TATA sequence.[12] Similarly, in humans only 24% of genes have promoter regions containing the TATA box.[13] Genes containing the TATA-box tend to be involved in stress-responses and certain types of metabolism and are more highly regulated when compared to TATA-less genes.[12][14] Generally, TATA-containing genes are not involved in essential cellular functions such as cell growth, DNA replication, transcription, and translation because of their highly regulated nature.[14]
The TATA box is usually located 25-35 base pairs upstream of the transcription start site. Genes containing the TATA box usually require additional promoter elements, including an initiator site located just upstream of the transcription start site and a downstream core element (DCE).[3] These additional promoter regions work in conjunction with the TATA box to regulate initiation of transcription in eukaryotes.
Function
editRole in transcription initiation
editThe TATA-box is the site of preinitiation complex formation, which is the first step in transcription initiation in eukaryotes. Formation of the preinitiation complex begins when the multi-subunit transcription factor II D (TFIID) binds to the TATA box at its TATA-binding protein (TBP) subunit.[3] TBP binds to the minor groove[15] of the TATA box via a region of antiparallel β sheets in the protein.[16] Three types of molecular interactions contribute to TBP binding to the TATA box:
- Four phenylalanine residues(Phe57, Phe74, Phe148, Phe 165) on TBP bind to DNA and form kinks in the DNA, forcing the DNA minor groove open.[16][17][18]
- Four hydrogen bonds form between polar side chains on TBP amino acid (Asn27, Asn117, Thr82, Thr173)( and bases in the minor groove.[16]
- Numerous hydrophobic interactions(~15) form between TBP residues(notably Ile152 and Leu163) and DNA bases, including van der Waals forces.[16][17][18]
Additionally, binding of TBP is facilitated by stabilizing interactions with DNA flanking the TATA box, which consists of G-C rich sequences.[19] These secondary interactions induce bending of the DNA and helical unwinding.[20] The degree of DNA bending is species and sequence dependent. For example, one study used the adenovirus TATA promoter sequence (5'-CGCTATAAAAGGGC-3') as a model binding sequence and found that human TBP binding to the TATA box induced a 97° bend toward the major groove while the yeast TBP protein only induced an 82° bend.[21] X-ray crystallography studies of TBP/TATA-box complexes generally agree that the DNA goes through an ~80° bend during the process of TBP-binding.[16][17][18]
The conformational changes induced by TBP binding to the TATA box allows for additional transcription factors and RNA polymerase II to bind to the promoter region. TFIID first binds to the TATA box, facilitated by TFIIA binding to the upstream part of the TFIID complex.[22][23] TFIIB then binds to the TFIID-TFIIA-DNA complex through interactions both upstream and downstream of the TATA box.[24] RNA polymerase II is then recruited to this multi-protein complex with the help of TFIIF.[24] Additional transcription factors then bind, first TFIIE and then TFIIH.[24] This completes the assembly of the preinitiation complex for eukaryotic transcription.[3] Generally, the TATA box is found at RNA polymerase II promoter regions, although some in vitro studies have demonstrated that RNA polymerase III can recognize TATA sequences.[25]
This cluster of RNA polymerase II and various transcription factors is known as the basal transcriptional complex (BTC). In this state, it only gives a low level of transcription. Other factors must stimulate the BTC to increase transcription levels.[2] One such example of a BTC stimulating region of DNA is the CAAT box. Additional factors, including the Mediator complex, transcriptional regulatory proteins, and nucleosome-modifying enzymes also enhance transcription in vivo.[3]
Interactions
editIn specific cell types or on specific promoters TBP can be replaced by one of several TBP-related factors (TRF1 in Drosophila, TBPL1/TRF2 in metazoans, TBPL2/TRF3 in vertebrates), some of which interact with the TATA box similar to TBP.[26] Interaction of TATA boxes with a variety of activators or repressors can influence the transcription of genes in many ways[citation needed]. Enhancers are long-range regulatory elements that increase promoter activity while silencers repress promoter activity.
Mutations
editMutations to the TATA box can range from a deletion or insertion to a point mutation with varying effects based on the gene that has been mutated. The mutations change the binding of the TATA-binding protein (TBP) for transcription initiation. Thus, there is a resulting change in phenotype based on the gene that is not being expressed (Figure 3).
Insertions or deletions
editOne of the first studies of TATA box mutations looked at a sequence of DNA from Agrobacterium tumefaciens for the octopine type cytokinin gene.[27] This specific gene has three TATA boxes. A phenotype change was only observed when all three TATA boxes were deleted. An insertion of extra base pairs between the last TATA box and the transcription start site resulted in a shift in the start site; thus, resulting in a phenotypic change. From this original mutation study, a change in transcription can be seen when there is no TATA box to promote transcription, but transcription of a gene will occur when there is an insertion to the sequence. The nature of the resulting phenotype may be affected due to the insertion.
Mutations in maize promoters affect the expression of the promoter genes in a plant-organ-specific manner.[29] A duplication of the TATA box leads to a significant decrease in enzymatic activity in the scutellum and roots, leaving pollen enzymatic levels unaffected. A deletion of the TATA box leads to a small decrease in enzymatic activity in the scutellum and roots, but a large decrease in enzymatic levels in pollen.[29]
Point mutations
editPoint mutations to the TATA box have similar varying phenotypic changes depending on the gene that is being affected. Studies also show that the placement of the mutation in the TATA box sequence hinders the binding of TBP.[28] For example, a mutation from TATAAAA to CATAAAA does completely hinder the binding sufficiently to change transcription, the neighboring sequences can affect if there is a change or not.[30] However, a change can be seen in HeLa cells with a TATAAAA to TATACAA which leads to a 20 fold decrease in transcription.[31] Some diseases that can be caused due to this insufficiency by specific gene transcription are: Thalassemia,[32] lung cancer,[33] chronic hemolytic anemia,[34] immunosuppression,[35] hemophilia B Leyden,[36] and thrombophlebitis and myocardial infarction.[37]
Savinkova et al. has written a simulation to predict the KD value for a selected TATA box sequence and TBP.[38] This can be used to directly predict the phenotypic traits resulting from a selected mutation based on how tightly TBP is binding to the TATA box.
Diseases
editMutations in the TATA box region affects the binding of the TATA-binding protein (TBP) for transcription initiation, which may cause carriers to have a disease phenotype.
Gastric cancer is correlated with TATA box polymorphism.[39] The TATA box has a binding site for the transcription factor of the PG2 gene. This gene produces PG2 serum, which is used as a biomarker for tumours in gastric cancer. Longer TATA box sequences correlates with higher levels of PG2 serum indicating gastric cancer conditions. Carriers with shorter TATA box sequences may produce lower levels of PG2 serum.
Several neurodegenerative disorders are associated TATA box mutations.[40] Two disorders have been highlighted, spinocerebellar ataxia and Huntington's disease. In spinocerebellar ataxia, the disease phenotype is caused by expansion of the polyglutamine repeat in the TATA-binding protein (TBP). An accumulation of these polyglutamine-TBP cells will occur, as shown by protein aggregates in brain sections of patients, resulting in a loss of neuronal cells.
Blindness can be caused by excessive cataract formation when the TATA box is targeted by microRNAs to increase the level of oxidative stress genes.[41] MicroRNAs can target the 3'-untranslated region and bind to the TATA box to activate the transcription of oxidative stress related genes.
SNPs in TATA boxes are associated with B-thalassemia, immunosuppression, and other neurological disorders.[42] SNPs destabilize the TBP/TATA complex which significantly decreases the rate at which TATA-binding proteins (TBP) will bind to the TATA box. This leads to lower levels of transcription affecting the severity of the disease. Results from studies have shown the interaction in vitro so far, but results may be comparable to that in vivo.
Gilbert's syndrome is correlated with UTG1A1 TATA box polymorphism.[43] This poses a risk for developing jaundice in newborns.
MicroRNAs also play a role in replicating viruses such as HIV-1.[44] Novel HIV-1-encoded microRNA have been found to enhance the production of the virus as well as activating HIV-1 latency by targeting the TATA box region.
Clinical significance
editTechnology
editMany of the studies so far have been performed in vitro, providing only a prediction of what may happen not a real-time representation of what is happening in the cells. Recent studies in 2016 have been done to demonstrate TATA-binding activity in vivo. Core promoter-specific mechanisms for transcription initiation by the canonical TBP/TFIID-dependent basal transcription machinery has recently been documented in vivo showing the activation by SRF-dependent upstream activating sequence (UAS) of the human ACTB gene involved in TATA-binding.[5]
Cancer therapy
editPharmaceutical companies have been designing cancer therapy drugs to target DNA in traditional methods over the years, and have proven to be successful.[45] However, the toxicity of these drugs have pushed scientists to explore other processes related to DNA that could be targeted instead. In recent years, a collective effort has been made to find cancer-specific molecular targets, such as protein-DNA complexes, which include the TATA binding motif. Compounds that trap the protein-DNA intermediate could result in it being toxic to the cell once they encounter a DNA processing event. Example of drugs that contain such compounds include topotecan, SN-38 (topoisomerase I), doxorubicin, and mitoxantrone (topoisomerase II).[45] Cisplatin is a compound that binds covalently to adjacent guanines in the major groove of DNA, which distorts DNA to allow access of DNA-binding proteins in the minor groove.[45] This will destabilize the interaction between the TATA-binding protein (TBP) to the TATA box. The result is to immobilize the TATA-binding protein (TBP) on DNA in order to down-regulate transcription initiation.
Genetic engineering
editTATA box modification
editEvolutionary changes have pushed plants to adapt to the changing environmental conditions. In the history of Earth, the development of Earth's aerobic atmosphere resulted in an iron deficiency in plants.[46] Compared to other members of the same species, Malus baccata var. xiaojinensis has a TATA box inserted in the promoter upstream of the iron-regulated transporter 1 (IRT1) promoter. As a result, the promoter activity levels are enhanced, increasing TFIID activity and subsequently transcription initiation, resulting in a more iron-efficient phenotype.[46]
See also
editReferences
edit- ^ a b c Lifton RP, Goldberg ML, Karp RW, Hogness DS (1978). "The organization of the histone genes in Drosophila melanogaster: functional and evolutionary implications". Cold Spring Harbor Symposia on Quantitative Biology. 42 (2): 1047–51. doi:10.1101/sqb.1978.042.01.105. PMID 98262.
- ^ a b c d e f Smale ST, Kadonaga JT (2003). "The RNA polymerase II core promoter". Annual Review of Biochemistry. 72: 449–79. doi:10.1146/annurev.biochem.72.121801.161520. PMID 12651739.
- ^ a b c d e f Watson, James D. (2014). Molecular biology of the gene. Watson, James D., 1928- (Seventh ed.). Boston. ISBN 9780321762436. OCLC 824087979.
{{cite book}}: CS1 maint: location missing publisher (link) - ^ a b Ohshima Y, Okada N, Tani T, Itoh Y, Itoh M (October 1981). "Nucleotide sequences of mouse genomic loci including a gene or pseudogene for U6 (4.8S) nuclear RNA". Nucleic Acids Research. 9 (19): 5145–58. doi:10.1093/nar/9.19.5145. PMC 327505. PMID 6171774.
- ^ a b c d Xu M, Gonzalez-Hurtado E, Martinez E (April 2016). "Core promoter-specific gene regulation: TATA box selectivity and Initiator-dependent bi-directionality of serum response factor-activated transcription". Biochimica et Biophysica Acta (BBA) - Gene Regulatory Mechanisms. 1859 (4): 553–63. doi:10.1016/j.bbagrm.2016.01.005. PMC 4818687. PMID 26824723.
- ^ Mainz D, Quadt I, Stranzenbach AK, Voss D, Guarino LA, Knebel-Mörsdorf D (June 2014). "Expression and nuclear localization of the TATA-box-binding protein during baculovirus infection". The Journal of General Virology. 95 (Pt 6): 1396–407. doi:10.1099/vir.0.059949-0. PMID 24676420. S2CID 33480957.
- ^ Gehring WJ (1998). Master Control Genes in Development and Evolution: The Homeobox Story. New Haven: Yale University Press. ISBN 978-0300074093.
- ^ a b Kutach AK, Kadonaga JT (July 2000). "The downstream promoter element DPE appears to be as widely used as the TATA box in Drosophila core promoters". Molecular and Cellular Biology. 20 (13): 4754–64. doi:10.1128/mcb.20.13.4754-4764.2000. PMC 85905. PMID 10848601.
- ^ Suzuki Y, Tsunoda T, Sese J, Taira H, Mizushima-Sugano J, Hata H, Ota T, Isogai T, Tanaka T, Nakamura Y, Suyama A, Sakaki Y, Morishita S, Okubo K, Sugano S (May 2001). "Identification and characterization of the potential promoter regions of 1031 kinds of human genes". Genome Research. 11 (5): 677–84. doi:10.1101/gr.gr-1640r. PMC 311086. PMID 11337467.
- ^ a b c Tripathi, G. (2010). Cellular and Biochemical Science. New Delhi: I.K. International Publishing House Pvt. Ltd. pp. 373–374. ISBN 9788188237852.
- ^ a b Baptista T, Grünberg S, Minoungou N, Koster MJ, Timmers HT, Hahn S, Devys D, Tora L (October 2017). "SAGA Is a General Cofactor for RNA Polymerase II Transcription". Molecular Cell. 68 (1): 130–143.e5. doi:10.1016/j.molcel.2017.08.016. PMC 5632562. PMID 28918903.
- ^ a b Basehoar AD, Zanton SJ, Pugh BF (March 2004). "Identification and distinct regulation of yeast TATA box-containing genes". Cell. 116 (5): 699–709. doi:10.1016/s0092-8674(04)00205-3. PMID 15006352.
- ^ Yang C, Bolotin E, Jiang T, Sladek FM, Martinez E (March 2007). "Prevalence of the initiator over the TATA box in human and yeast genes and identification of DNA motifs enriched in human TATA-less core promoters". Gene. 389 (1): 52–65. doi:10.1016/j.gene.2006.09.029. PMC 1955227. PMID 17123746.
- ^ a b Bae SH, Han HW, Moon J (2015). "Functional analysis of the molecular interactions of TATA box-containing genes and essential genes". PLOS ONE. 10 (3): e0120848. Bibcode:2015PLoSO..1020848B. doi:10.1371/journal.pone.0120848. PMC 4366266. PMID 25789484.
- ^ Starr DB, Hawley DK (December 1991). "TFIID binds in the minor groove of the TATA box". Cell. 67 (6): 1231–40. doi:10.1016/0092-8674(91)90299-e. PMID 1760847. S2CID 10297041.
- ^ a b c d e Kim JL, Nikolov DB, Burley SK (October 1993). "Co-crystal structure of TBP recognizing the minor groove of a TATA element". Nature. 365 (6446): 520–7. Bibcode:1993Natur.365..520K. doi:10.1038/365520a0. PMID 8413605. S2CID 4371241.
- ^ a b c Nikolov DB, Chen H, Halay ED, Hoffman A, Roeder RG, Burley SK (May 1996). "Crystal structure of a human TATA box-binding protein/TATA element complex". Proceedings of the National Academy of Sciences of the United States of America. 93 (10): 4862–7. Bibcode:1996PNAS...93.4862N. doi:10.1073/pnas.93.10.4862. PMC 39370. PMID 8643494.
- ^ a b c Kim Y, Geiger JH, Hahn S, Sigler PB (October 1993). "Crystal structure of a yeast TBP/TATA-box complex". Nature. 365 (6446): 512–20. Bibcode:1993Natur.365..512K. doi:10.1038/365512a0. PMID 8413604. S2CID 4336203.
- ^ Horikoshi M, Bertuccioli C, Takada R, Wang J, Yamamoto T, Roeder RG (February 1992). "Transcription factor TFIID induces DNA bending upon binding to the TATA element". Proceedings of the National Academy of Sciences of the United States of America. 89 (3): 1060–4. Bibcode:1992PNAS...89.1060H. doi:10.1073/pnas.89.3.1060. PMC 48385. PMID 1736286.
- ^ Blair RH, Goodrich JA, Kugel JF (September 2012). "Single-molecule fluorescence resonance energy transfer shows uniformity in TATA binding protein-induced DNA bending and heterogeneity in bending kinetics". Biochemistry. 51 (38): 7444–55. doi:10.1021/bi300491j. PMC 3551999. PMID 22934924.
- ^ Whittington JE, Delgadillo RF, Attebury TJ, Parkhurst LK, Daugherty MA, Parkhurst LJ (July 2008). "TATA-binding protein recognition and bending of a consensus promoter are protein species dependent". Biochemistry. 47 (27): 7264–73. doi:10.1021/bi800139w. PMID 18553934. S2CID 7460689.
- ^ Louder RK, He Y, López-Blanco JR, Fang J, Chacón P, Nogales E (March 2016). "Structure of promoter-bound TFIID and model of human pre-initiation complex assembly". Nature. 531 (7596): 604–9. Bibcode:2016Natur.531..604L. doi:10.1038/nature17394. PMC 4856295. PMID 27007846.
- ^ Wang J, Zhao S, He W, Wei Y, Zhang Y, Pegg H, Shore P, Roberts SG, Deng W (July 2017). "A transcription factor IIA-binding site differentially regulates RNA polymerase II-mediated transcription in a promoter context-dependent manner". The Journal of Biological Chemistry. 292 (28): 11873–11885. doi:10.1074/jbc.M116.770412. PMC 5512080. PMID 28539359.
- ^ a b c Krishnamurthy S, Hampsey M (February 2009). "Eukaryotic transcription initiation". Current Biology. 19 (4): R153–6. doi:10.1016/j.cub.2008.11.052. PMID 19243687.
- ^ Duttke SH (July 2014). "RNA polymerase III accurately initiates transcription from RNA polymerase II promoters in vitro". The Journal of Biological Chemistry. 289 (29): 20396–404. doi:10.1074/jbc.M114.563254. PMC 4106352. PMID 24917680.
- ^ Akhtar W, Veenstra GJ (1 January 2011). "TBP-related factors: a paradigm of diversity in transcription initiation". Cell & Bioscience. 1 (1): 23. doi:10.1186/2045-3701-1-23. PMC 3142196. PMID 21711503.
- ^ a b Chioin R, Stritoni P, Scognamiglio R, Boffa GM, Daliento L, Razzolini R, Ramondo A, Dalla Volta S (1987). "[Natural history of coronary disease with and without aortocoronary by-pass operation. Survival curves of 272 patients over a maximum period of 24 months (author's transl)]". Giornale Italiano di Cardiologia. 8 (4): 359–64. doi:10.1093/nar/15.20.8283. PMC 306359. PMID 3671084.
- ^ a b Gaillard J, Haguenauer JP, Romanet P, Boulud B, Gerard JP (November 1977). "[Tumors of the olfactory placode. Study of 5 cases]". Journal Français d'Oto-Rhino-Laryngologie; Audiophonologie, Chirurgie Maxillo-Faciale. 26 (9): 669–76. doi:10.1093/nar/24.15.3100. PMC 146060. PMID 8760900.
- ^ a b Kloeckener-Gruissem B, Vogel JM, Freeling M (January 1992). "The TATA box promoter region of maize Adh1 affects its organ-specific expression". The EMBO Journal. 11 (1): 157–66. doi:10.1002/j.1460-2075.1992.tb05038.x. PMC 556436. PMID 1740103.
- ^ Fei YJ, Stoming TA, Efremov GD, Efremov DG, Battacharia R, Gonzalez-Redondo JM, Altay C, Gurgey A, Huisman TH (June 1988). "Beta-thalassemia due to a T----A mutation within the ATA box". Biochemical and Biophysical Research Communications. 153 (2): 741–7. doi:10.1016/S0006-291X(88)81157-4. PMID 3382401.
- ^ Bower GC (1978). "The award of the Will Ross Medal for 1978". The American Review of Respiratory Disease. 118 (3): 635–636. PMID 360896.
- ^ Antonarakis SE, Irkin SH, Cheng TC, Scott AF, Sexton JP, Trusko SP, Charache S, Kazazian HH (1984). "beta-Thalassemia in American Blacks: novel mutations in the "TATA" box and an acceptor splice site". Proceedings of the National Academy of Sciences of the United States of America. 81 (4): 1154–8. Bibcode:1984PNAS...81.1154A. doi:10.1073/pnas.81.4.1154. PMC 344784. PMID 6583702.
- ^ Zienolddiny S, Ryberg D, Maggini V, Skaug V, Canzian F, Haugen A (April 2004). "Polymorphisms of the interleukin-1 beta gene are associated with increased risk of non-small cell lung cancer". International Journal of Cancer. 109 (3): 353–6. doi:10.1002/ijc.11695. PMID 14961572.
- ^ Hildebrandt P (August 1991). "Subcutaneous absorption of insulin in insulin-dependent diabetic patients. Influence of species, physico-chemical properties of insulin and physiological factors". Danish Medical Bulletin. 38 (4): 337–46. PMC 1914533. PMID 8571957.
- ^ Takahashi K, Ezekowitz RA (November 2005). "The role of the mannose-binding lectin in innate immunity". Clinical Infectious Diseases. 41 (Suppl 7): S440–4. doi:10.1086/431987. PMID 16237644.
- ^ Sweet D, Golomb H, Desser R, Ultmann JE, Yachnin S, Stein R (May 1975). "Letter: Chemotherapy of advanced histocytic lymphomas". Lancet. 1 (7916): 6300–3. doi:10.1016/s0140-6736(75)92521-0. PMC 49488. PMID 1631121.
- ^ Arnaud E, Barbalat V, Nicaud V, Cambien F, Evans A, Morrison C, Arveiler D, Luc G, Ruidavets JB, Emmerich J, Fiessinger JN, Aiach M (March 2000). "Polymorphisms in the 5' regulatory region of the tissue factor gene and the risk of myocardial infarction and venous thromboembolism: the ECTIM and PATHROS studies. Etude Cas-Témoins de l'Infarctus du Myocarde. Paris Thrombosis case-control Study". Arteriosclerosis, Thrombosis, and Vascular Biology. 20 (3): 892–8. doi:10.1161/01.ATV.20.3.892. PMID 10712418.
- ^ Savinkova L, Drachkova I, Arshinova T, Ponomarenko P, Ponomarenko M, Kolchanov N (2013). "An experimental verification of the predicted effects of promoter TATA-box polymorphisms associated with human diseases on interactions between the TATA boxes and TATA-binding protein". PLOS ONE. 8 (2): e54626. Bibcode:2013PLoSO...854626S. doi:10.1371/journal.pone.0054626. PMC 3570547. PMID 23424617.
- ^ De Re V, Magris R, De Zorzi M, Maiero S, Caggiari L, Fornasarig M, Repetto O, Buscarini E, Di Mario F (2017). "P.08.10: Interference of PG2 Tata Box Region with the Serum PG2 Level in Gastric Cancer". Digestive and Liver Disease. 49: e182–e183. doi:10.1016/s1590-8658(17)30534-0. S2CID 79101992.
- ^ Roshan R, Choudhary A, Bhambri A, Bakshi B, Ghosh T, Pillai B (August 2017). "microRNA dysregulation in polyglutamine toxicity of TATA-box binding protein is mediated through STAT1 in mouse neuronal cells". Journal of Neuroinflammation. 14 (1): 155. doi:10.1186/s12974-017-0925-3. PMC 5543588. PMID 28774347.
- ^ Wu C, Liu Z, Ma L, Pei C, Qin L, Gao N, Li J, Yin Y (August 2017). "MiRNAs regulate oxidative stress related genes via binding to the 3' UTR and TATA-box regions: a new hypothesis for cataract pathogenesis". BMC Ophthalmology. 17 (1): 142. doi:10.1186/s12886-017-0537-9. PMC 5556341. PMID 28806956.
- ^ Drachkova I, Savinkova L, Arshinova T, Ponomarenko M, Peltek S, Kolchanov N (May 2014). "The mechanism by which TATA-box polymorphisms associated with human hereditary diseases influence interactions with the TATA-binding protein". Human Mutation. 35 (5): 601–8. doi:10.1002/humu.22535. PMID 24616209. S2CID 19928327.
- ^ Žaja O, Tiljak MK, Štefanović M, Tumbri J, Jurčić Z (May 2014). "Correlation of UGT1A1 TATA-box polymorphism and jaundice in breastfed newborns-early presentation of Gilbert's syndrome". The Journal of Maternal-Fetal & Neonatal Medicine. 27 (8): 844–50. doi:10.3109/14767058.2013.837879. PMID 23981182. S2CID 29893463.
- ^ Zhang Y, Fan M, Geng G, Liu B, Huang Z, Luo H, Zhou J, Guo X, Cai W, Zhang H (March 2014). "A novel HIV-1-encoded microRNA enhances its viral replication by targeting the TATA box region". Retrovirology. 11: 23. doi:10.1186/1742-4690-11-23. PMC 4007588. PMID 24620741.
- ^ a b c Hurley LH (March 2002). "DNA and its associated processes as targets for cancer therapy". Nature Reviews. Cancer. 2 (3): 188–200. doi:10.1038/nrc749. PMID 11990855. S2CID 24209612.
- ^ a b Zhang M, Lv Y, Wang Y, Rose JK, Shen F, Han Z, Zhang X, Xu X, Wu T, Han Z (January 2017). "TATA Box Insertion Provides a Selection Mechanism Underpinning Adaptations to Fe Deficiency". Plant Physiology. 173 (1): 715–727. doi:10.1104/pp.16.01504. PMC 5210749. PMID 27881725.