Influenza hemagglutinin (HA) or haemagglutinin[p] (British English) is a homotrimeric glycoprotein found on the surface of influenza viruses and is integral to its infectivity.
| Hemagglutinin | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 | |||||||||
| Identifiers | |||||||||
| Symbol | Hemagglutinin | ||||||||
| Pfam | PF00509 | ||||||||
| InterPro | IPR001364 | ||||||||
| SCOP2 | 1hgd / SCOPe / SUPFAM | ||||||||
| OPM superfamily | 109 | ||||||||
| OPM protein | 6hjq | ||||||||
| |||||||||
| Influenza C hemagglutinin stalk | |||||||||
|---|---|---|---|---|---|---|---|---|---|
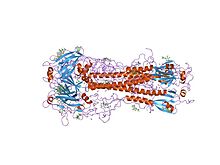
 x-ray structure of the haemagglutinin-esterase-fusion glycoprotein of influenza c virus | |||||||||
| Identifiers | |||||||||
| Symbol | Hema_stalk | ||||||||
| Pfam | PF08720 | ||||||||
| InterPro | IPR014831 | ||||||||
| SCOP2 | 1flc / SCOPe / SUPFAM | ||||||||
| OPM superfamily | 277 | ||||||||
| OPM protein | 2jrd | ||||||||
| |||||||||
Hemagglutinin is a class I fusion protein,[1][2] having multifunctional activity as both an attachment factor and membrane fusion protein. Therefore, HA is responsible for binding influenza viruses to sialic acid on the surface of target cells, such as cells in the upper respiratory tract or erythrocytes,[3] resulting in the internalization of the virus.[4] Additionally, HA is responsible for the fusion of the viral envelope with the late endosomal membrane once exposed to low pH (5.0–5.5).[5]
The name "hemagglutinin" comes from the protein's ability to cause red blood cells (i.e., erythrocytes) to clump together (i.e., agglutinate) in vitro.[6]
Subtypes
editHemagglutinin (HA) in influenza A virus (IAV) has at least 18 different subtypes.[7] These subtypes are named H1 through H18. H16 was discovered in 2004 on IAVs isolated from black-headed gulls from Sweden and Norway. H17 was discovered in 2012 in fruit bats.[8][9] Most recently, H18 was discovered in a Peruvian bat in 2013.[10] The first three hemagglutinins, H1, H2, and H3, are found in influenza viruses that infect humans. By phylogenetic similarity, the HA proteins are divided into 2 groups, with H1, H2, H5, H6, H8, H9, H11, H12, H13, H16, H17, and H18 belonging to group 1 and the rest in group 2.[11] The serotype of IAV is determined by the HA and neuraminidase (NA) proteins expressed on its surface.[12] Neuraminidase has 11 known subtypes; hence, influenza viruses are named according to the combinations of HA and NA proteins expressed (e.g., H1N1 and H5N2).[7]
A highly pathogenic avian influenza A virus, A(H5N1), is known to infect humans as well as its original avian hosts, albeit quite infrequently.[11] It has been reported that single amino acid changes in the virus's H5 hemagglutinin have been found in human patients that "can significantly alter receptor specificity of avian H5N1 viruses, providing them with an ability to bind to receptors optimal for human influenza viruses."[13][14] This finding seems to explain how an H5N1 virus that normally does not infect humans can mutate and become able to efficiently infect human cells. The hemagglutinin of the H5N1 virus has been associated with its high degree of pathogenicity, apparently due to its ease of conversion to an active form by proteolysis.[15][16]
Structure
editHA is a homotrimeric integral membrane glycoprotein. It has C3 molecular symmetry. It is shaped like a cylinder, and is approximately 13.5 nanometres long.[17][18] HA trimer is made of three identical monomers. Each monomer is made of an intact HA0 single polypeptide chain with HA1 and HA2 regions that are linked by 2 disulfide bridges.[18][19] Each HA2 region adopts alpha helical coiled coil structure and sits on top of the HA1 region, which is a small globular domain that consists of a mix of α/β structures.[20] The HA trimer is synthesized as inactive precursor protein HA0 to prevent any premature and unwanted fusion activity and must be cleaved by host proteases in order to be infectious. At neutral pH, the 23 residues near the N-terminus of HA2, also known as the fusion peptide that is eventually responsible for fusion between viral and host membrane, is hidden in a hydrophobic pocket between the HA2 trimeric interface.[21] The C-terminus of HA2, also known as the transmembrane domain, spans the viral membrane and anchors protein to the membrane.[22]
- HA1
- HA1 is mostly composed of antiparallel beta-sheets.[17]
- HA2
- HA2 domain contains three long alpha helices, one from each monomer. Each of these helices is connected by a flexible, loop region called Loop-B (residue 59 to 76).[23]
Function
editHA plays two key functions in viral entry. Firstly, it allows the recognition of target vertebrate cells, accomplished through the binding to these cells' sialic acid-containing receptors. Secondly, once bound it facilitates the entry of the viral genome into the target cells by causing the fusion of host endosomal membrane with the viral membrane.[24]
Specifically, the HA1 domain of the protein binds to the monosaccharide sialic acid which is present on the surface of its target cells, allowing attachment of viral particle to the host cell surface. HA17 and HA18 have been described to bind MHC class II molecules as a receptor for entry rather than sialic acid.[25] The host cell membrane then engulfs the virus, a process known as endocytosis, and pinches off to form a new membrane-bound compartment within the cell called an endosome. The cell then attempts to begin digesting the contents of the endosome by acidifying its interior and transforming it into a lysosome. Once the pH within the endosome drops to about 5.0 to 6.0, a series of conformational rearrangement occurs to the protein. First, fusion peptide is released from the hydrophobic pocket and HA1 is dissociated from HA2 domain. HA2 domain then undergoes extensive conformation change that eventually bring the two membranes into close contact.[citation needed]
This so-called "fusion peptide" that was released as pH is lowered, acts like a molecular grappling hook by inserting itself into the endosomal membrane and locking on. Then, HA2 refolds into a new structure (which is more stable at the lower pH), it "retracts the grappling hook" and pulls the endosomal membrane right up next to the virus particle's own membrane, causing the two to fuse together. Once this has happened, the contents of the virus such as viral RNA are released in the host cell's cytoplasm and then transported to the host cell nucleus for replication.[26]
As a treatment target
editSince hemagglutinin is the major surface protein of the influenza A virus and is essential to the entry process, it is the primary target of neutralizing antibodies. [citation needed]These antibodies against flu have been found to act by two different mechanisms, mirroring the dual functions of hemagglutinin:
Head antibodies
editSome antibodies against hemagglutinin act by inhibiting attachment. This is because these antibodies bind near the top of the hemagglutinin "head" (blue region in figure above) and physically block the interaction with sialic acid receptors on target cells.[27]
Stem antibodies
editThis group of antibodies acts by preventing membrane fusion (only in vitro; the efficacy of these antibodies in vivo is believed to be a result of antibody-dependent cell-mediated cytotoxicity and the complement system).[28]
The stem or stalk region of HA (HA2), is highly conserved across different strains of influenza viruses. The conservation makes it an attractive target for broadly neutralizing antibodies that target all flu subtypes, and for developing universal vaccines that let humans produce these antibodies naturally.[29] Its structural changes from prefusion to postfusion conformation drives fusion between viral membrane and host membrane. Therefore, antibodies targeting this region can block key structural changes that eventually drive the membrane fusion process, and therefore are able to achieve antiviral activity against several influenza virus subtypes. At least one fusion-inhibiting antibody was found to bind closer to the top of hemagglutinin, and is thought to work by cross-linking the heads together, the opening of which is thought to be the first step in the membrane fusion process.[30]
Examples are human antibodies F10,[31] FI6,[32] CR6261. They recognize sites in the stem/stalk region (orange region in figure at right), far away from the receptor binding site.[33][34]
In 2015 researchers designed an immunogen mimicking the HA stem, specifically the area where the antibody ties to the virus of the antibody CR9114. Rodent and nonhuman primate models given the immunogen produced antibodies that could bind with HAs in many influenza subtypes, including H5N1.[35] When the HA head is present, the immune system does not generally make bNAbs (broadly neutralizing antibodies). Instead, it makes the head antibodies that only recognize a few subtypes. Since the head is responsible for holding the three HA units together, a stem-only HA needs its own way to hold itself together. One team designed self-assembling HA-stem nanoparticles, using a protein called ferritin to hold the HA together. Another replaced and added amino acids to stabilize a mini-HA lacking a proper head.[citation needed]
A 2016 vaccine trial in humans found many broadly neutralizing antibodies targeting the stem produced by the immune system. Three classes of highly similar antibodies were recovered from multiple human volunteers, suggesting that a universal vaccine that produces reproducible antibodies is indeed possible.[36]
Other agents
editThere are also other hemagglutinin-targeted influenza virus inhibitors that are not antibodies:[37]
- Arbidol
- Small Molecules
- Natural compounds
- Proteins and peptides
See also
editNotes
editReferences
edit- ^ Skehel JJ, Wiley DC (July 2000). "Receptor binding and membrane fusion in virus entry: the influenza hemagglutinin". Annual Review of Biochemistry. 69. Annual Reviews: 531–569. doi:10.1146/annurev.biochem.69.1.531. PMID 10966468.
- ^ Kielian M, Rey FA (January 2006). "Virus membrane-fusion proteins: more than one way to make a hairpin". Nature Reviews. Microbiology. 4 (1): 67–76. doi:10.1038/nrmicro1326. PMC 7097298. PMID 16357862.
- ^ Russell RJ, Kerry PS, Stevens DJ, Steinhauer DA, Martin SR, Gamblin SJ, et al. (November 2008). "Structure of influenza hemagglutinin in complex with an inhibitor of membrane fusion". Proceedings of the National Academy of Sciences of the United States of America. 105 (46): 17736–17741. Bibcode:2008PNAS..10517736R. doi:10.1073/pnas.0807142105. PMC 2584702. PMID 19004788.
- ^ Edinger TO, Pohl MO, Stertz S (February 2014). "Entry of influenza A virus: host factors and antiviral targets". The Journal of General Virology. 95 (Pt 2): 263–277. doi:10.1099/vir.0.059477-0. PMID 24225499.
- ^ Banerjee I, Yamauchi Y, Helenius A, Horvath P (12 July 2013). "High-content analysis of sequential events during the early phase of influenza A virus infection". PLOS ONE. 8 (7): e68450. Bibcode:2013PLoSO...868450B. doi:10.1371/journal.pone.0068450. PMC 3709902. PMID 23874633.
- ^ Nelson DL, Cox MM (2005). Lehninger's Principles of Biochemistry (4th ed.). New York: WH Freeman.
- ^ a b CDC (30 March 2023). "Types of Influenza Viruses". Centers for Disease Control and Prevention. Retrieved 12 May 2024.
- ^ Fouchier RA, Munster V, Wallensten A, Bestebroer TM, Herfst S, Smith D, et al. (March 2005). "Characterization of a novel influenza A virus hemagglutinin subtype (H16) obtained from black-headed gulls". Journal of Virology. 79 (5): 2814–22. doi:10.1128/JVI.79.5.2814-2822.2005. PMC 548452. PMID 15709000.
- ^ "Unique new flu virus found in bats". Archived from the original on 20 May 2012.
- ^ Tong S, Zhu X, Li Y, Shi M, Zhang J, Bourgeois M, et al. (October 2013). "New world bats harbor diverse influenza A viruses". PLOS Pathogens. 9 (10): e1003657. doi:10.1371/journal.ppat.1003657. PMC 3794996. PMID 24130481.
- ^ a b Sutton TC, Chakraborty S, Mallajosyula VV, Lamirande EW, Ganti K, Bock KW, et al. (15 December 2017). "Protective efficacy of influenza group 2 hemagglutinin stem-fragment immunogen vaccines". npj Vaccines. 2 (1): 35. doi:10.1038/s41541-017-0036-2. PMC 5732283. PMID 29263889.
- ^ "Influenza Type A Viruses". Avian Influenza (Flu). CDC. 19 April 2017. Retrieved 27 August 2018.
- ^ Suzuki Y (March 2005). "Sialobiology of influenza: molecular mechanism of host range variation of influenza viruses". Biological & Pharmaceutical Bulletin. 28 (3): 399–408. doi:10.1248/bpb.28.399. PMID 15744059.
- ^ Gambaryan A, Tuzikov A, Pazynina G, Bovin N, Balish A, Klimov A (January 2006). "Evolution of the receptor binding phenotype of influenza A (H5) viruses". Virology. 344 (2): 432–8. doi:10.1016/j.virol.2005.08.035. PMID 16226289.
- ^ Hatta M, Gao P, Halfmann P, Kawaoka Y (September 2001). "Molecular basis for high virulence of Hong Kong H5N1 influenza A viruses". Science. 293 (5536): 1840–2. Bibcode:2001Sci...293.1840H. doi:10.1126/science.1062882. PMID 11546875. S2CID 37415902.
- ^ Senne DA, Panigrahy B, Kawaoka Y, Pearson JE, Süss J, Lipkind M, et al. (1996). "Survey of the hemagglutinin (HA) cleavage site sequence of H5 and H7 avian influenza viruses: amino acid sequence at the HA cleavage site as a marker of pathogenicity potential". Avian Diseases. 40 (2): 425–37. doi:10.2307/1592241. JSTOR 1592241. PMID 8790895.
- ^ a b Wilson IA, Skehel JJ, Wiley DC (January 1981). "Structure of the haemagglutinin membrane glycoprotein of influenza virus at 3 A resolution". Nature. 289 (5796): 366–73. Bibcode:1981Natur.289..366W. doi:10.1038/289366a0. PMID 7464906. S2CID 4327688.
- ^ a b Boonstra S, Blijleven JS, Roos WH, Onck PR, van der Giessen E, van Oijen AM (May 2018). "Hemagglutinin-Mediated Membrane Fusion: A Biophysical Perspective" (PDF). Annual Review of Biophysics. 47 (1): 153–173. doi:10.1146/annurev-biophys-070317-033018. PMID 29494252.
- ^ Di Lella S, Herrmann A, Mair CM (June 2016). "Modulation of the pH Stability of Influenza Virus Hemagglutinin: A Host Cell Adaptation Strategy". Biophysical Journal. 110 (11): 2293–2301. Bibcode:2016BpJ...110.2293D. doi:10.1016/j.bpj.2016.04.035. PMC 4906160. PMID 27276248.
- ^ Smrt ST, Lorieau JL (2016). "Membrane Fusion and Infection of the Influenza Hemagglutinin". Protein Reviews. Advances in Experimental Medicine and Biology. Vol. 966. Springer Singapore. pp. 37–54. doi:10.1007/5584_2016_174. ISBN 9789811069215. PMID 27966108.
- ^ Wiley DC, Skehel JJ (June 1987). "The structure and function of the hemagglutinin membrane glycoprotein of influenza virus". Annual Review of Biochemistry. 56 (1): 365–94. doi:10.1146/annurev.bi.56.070187.002053. PMID 3304138.
- ^ Strauss JH, Strauss EG (2008). Viruses and human disease (2nd ed.). Amsterdam: Elsevier / Academic Press. ISBN 9780080553160. OCLC 630107686.
- ^ Stevens J, Corper AL, Basler CF, Taubenberger JK, Palese P, Wilson IA (March 2004). "Structure of the uncleaved human H1 hemagglutinin from the extinct 1918 influenza virus". Science. 303 (5665): 1866–70. Bibcode:2004Sci...303.1866S. doi:10.1126/science.1093373. PMID 14764887. S2CID 42010652.
- ^ White JM, Hoffman LR, Arevalo JH, et al. (1997). "Attachment and entry of influenza virus into host cells. Pivotal roles of hemagglutinin". In Chiu W, Burnett RM, Garcea RL (eds.). Structural Biology of Viruses. Oxford University Press. pp. 80–104.
- ^ Karakus U, Thamamongood T, Ciminski K, Ran W, Günther SC, Pohl MO, et al. (March 2019). "MHC class II proteins mediate cross-species entry of bat influenza viruses". Nature. 567 (7746): 109–112. Bibcode:2019Natur.567..109K. doi:10.1038/s41586-019-0955-3. PMID 30787439. S2CID 67769877.
- ^ Mair CM, Ludwig K, Herrmann A, Sieben C (April 2014). "Receptor binding and pH stability – how influenza A virus hemagglutinin affects host-specific virus infection". Biochimica et Biophysica Acta (BBA) - Biomembranes. 1838 (4): 1153–1168. doi:10.1016/j.bbamem.2013.10.004. PMID 24161712.
- ^ Goh BC, Rynkiewicz MJ, Cafarella TR, White MR, Hartshorn KL, Allen K, et al. (November 2013). "Molecular mechanisms of inhibition of influenza by surfactant protein D revealed by large-scale molecular dynamics simulation". Biochemistry. 52 (47): 8527–38. doi:10.1021/bi4010683. PMC 3927399. PMID 24224757.
- ^ DiLillo DJ, Tan GS, Palese P, Ravetch JV (February 2014). "Broadly neutralizing hemagglutinin stalk-specific antibodies require FcγR interactions for protection against influenza virus in vivo". Nature Medicine. 20 (2): 143–51. doi:10.1038/nm.3443. PMC 3966466. PMID 24412922.
- ^ Sautto GA, Kirchenbaum GA, Ross TM (January 2018). "Towards a universal influenza vaccine: different approaches for one goal". Virology Journal. 15 (1): 17. doi:10.1186/s12985-017-0918-y. PMC 5785881. PMID 29370862.
- ^ Barbey-Martin C, Gigant B, Bizebard T, Calder LJ, Wharton SA, Skehel JJ, et al. (March 2002). "An antibody that prevents the hemagglutinin low pH fusogenic transition". Virology. 294 (1): 70–4. doi:10.1006/viro.2001.1320. PMID 11886266.
- ^ Sui J, Hwang WC, Perez S, Wei G, Aird D, Chen LM, et al. (March 2009). "Structural and functional bases for broad-spectrum neutralization of avian and human influenza A viruses". Nature Structural & Molecular Biology. 16 (3): 265–73. doi:10.1038/nsmb.1566. PMC 2692245. PMID 19234466.
- ^ Corti D, Voss J, Gamblin SJ, Codoni G, Macagno A, Jarrossay D, et al. (August 2011). "A neutralizing antibody selected from plasma cells that binds to group 1 and group 2 influenza A hemagglutinins". Science. 333 (6044): 850–6. Bibcode:2011Sci...333..850C. doi:10.1126/science.1205669. PMID 21798894. S2CID 5086468.
- ^ Throsby M, van den Brink E, Jongeneelen M, Poon LL, Alard P, Cornelissen L, et al. (2008). "Heterosubtypic neutralizing monoclonal antibodies cross-protective against H5N1 and H1N1 recovered from human IgM+ memory B cells". PLOS ONE. 3 (12): e3942. Bibcode:2008PLoSO...3.3942T. doi:10.1371/journal.pone.0003942. PMC 2596486. PMID 19079604.
- ^ Ekiert DC, Bhabha G, Elsliger MA, Friesen RH, Jongeneelen M, Throsby M, et al. (April 2009). "Antibody recognition of a highly conserved influenza virus epitope". Science. 324 (5924): 246–51. Bibcode:2009Sci...324..246E. doi:10.1126/science.1171491. PMC 2758658. PMID 19251591.
- ^ Micu A (25 August 2015). "Universal flu vaccine: research moves closer". ZME Science. Retrieved 10 June 2016.
- ^ Joyce MG, Wheatley AK, Thomas PV, Chuang GY, Soto C, Bailer RT, et al. (July 2016). "Vaccine-Induced Antibodies that Neutralize Group 1 and Group 2 Influenza A Viruses". Cell. 166 (3): 609–623. doi:10.1016/j.cell.2016.06.043. PMC 4978566. PMID 27453470.
- ^ Zeng LY, Yang J, Liu S (January 2017). "Investigational hemagglutinin-targeted influenza virus inhibitors". Expert Opinion on Investigational Drugs. 26 (1): 63–73. doi:10.1080/13543784.2017.1269170. PMID 27918208. S2CID 11401818.
- ^ Boyd RS (24 May 2007) [6 October 2005]. "Scientists race to develop a vaccine against a killer flu". Knight Ridder Newspapers. Retrieved 24 May 2018 – via Mcclatchydc.com.
- ^ "Bird flu: Don't fly into a panic – Harvard Health". Harvard.edu. October 2006. Retrieved 24 May 2018.
External links
edit- Jmol tutorial of influenza hemagglutinin structure and activity.
- PDB Molecule of the Month Hemagglutinin (April 2006)
- Influenza Research Database Database of influenza protein sequences and structures
- 3D macromolecular structures of influenza hemagglutinin from the EM Data Bank(EMDB)
- Overview of all the structural information available in the PDB for UniProt: K7N5L2 (Influenza A virus Hemagglutinin) at the PDBe-KB.
- Overview of all the structural information available in the PDB for UniProt: A0A0J9X268 (H6N1 subtype Hemagglutinin) at the PDBe-KB.
- Overview of all the structural information available in the PDB for UniProt: A0A0J9X267 (H6N1 subtype Hemagglutinin HA2 chain) at the PDBe-KB.